A Flavone Synthase that Alters Lignin
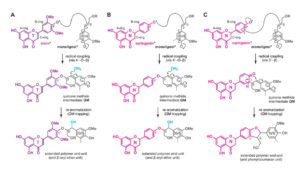 Lignin, a ubiquitous phenylpropanoid polymer found in the cell walls of vascular plants, is derived primarily from oxidative couplings of monolignols (p-hydroxycinnamyl alcohols). By filling up spaces between cell wall polysaccharides (cellulose and hemicelluloses), lignin confers increased mechanical strength, imperviousness, and resistance to pathogens. Lignin biosynthesis bioengineering has long been a major research, particularly because of its economic importance associated with agroindustrial uses of biomass. Lignin has traditionally been viewed as an impediment to chemical pulping, forage digestion by livestock, and cellulosic bioethanol production. Recently, it was discovered that many grasses, including cereals, utilize a member of the flavonoids, tricin (39,59-dimethoxyflavone), as a natural co-monomer with monolignols for cell wall lignification. Cytochrome P450 93G1 is a flavone synthase II (OsFNSII) indispensable for the biosynthesis of soluble tricin-derived metabolites in rice (Oryza sativa). Lam et al. () have examined the involvement of OsFNSII in lignification and alterations in cell wall properties upon tricin deficiency. A rice fnsII mutant was subjected to a series of analyses for the assessment of growth phenotypes, gene expression, as well as lignin structure. The mutant is similar in growth to wild-type control plants with normal vascular morphology. Chemical and nuclear magnetic resonance structural analyses demonstrated that the mutant lignin is completely devoid of tricin, indicating that FNSII activity is essential for the deposition of tricin-bound lignin in rice cell walls. The mutant also showed substantially reduced lignin content with decreased syringyl/guaiacyl lignin unit composition. Interestingly, the loss of tricin in the mutant lignin appears to be partially compensated by incorporating naringenin, which is a preferred substrate of OsFNSII. Such lignin alterations resulted in enhanced cell wall digestibility without negative impact on growth and development. Thus, grass biomass utilization may potentially be enhanced by manipulation of the flavone biosynthesis pathway.
Lignin, a ubiquitous phenylpropanoid polymer found in the cell walls of vascular plants, is derived primarily from oxidative couplings of monolignols (p-hydroxycinnamyl alcohols). By filling up spaces between cell wall polysaccharides (cellulose and hemicelluloses), lignin confers increased mechanical strength, imperviousness, and resistance to pathogens. Lignin biosynthesis bioengineering has long been a major research, particularly because of its economic importance associated with agroindustrial uses of biomass. Lignin has traditionally been viewed as an impediment to chemical pulping, forage digestion by livestock, and cellulosic bioethanol production. Recently, it was discovered that many grasses, including cereals, utilize a member of the flavonoids, tricin (39,59-dimethoxyflavone), as a natural co-monomer with monolignols for cell wall lignification. Cytochrome P450 93G1 is a flavone synthase II (OsFNSII) indispensable for the biosynthesis of soluble tricin-derived metabolites in rice (Oryza sativa). Lam et al. () have examined the involvement of OsFNSII in lignification and alterations in cell wall properties upon tricin deficiency. A rice fnsII mutant was subjected to a series of analyses for the assessment of growth phenotypes, gene expression, as well as lignin structure. The mutant is similar in growth to wild-type control plants with normal vascular morphology. Chemical and nuclear magnetic resonance structural analyses demonstrated that the mutant lignin is completely devoid of tricin, indicating that FNSII activity is essential for the deposition of tricin-bound lignin in rice cell walls. The mutant also showed substantially reduced lignin content with decreased syringyl/guaiacyl lignin unit composition. Interestingly, the loss of tricin in the mutant lignin appears to be partially compensated by incorporating naringenin, which is a preferred substrate of OsFNSII. Such lignin alterations resulted in enhanced cell wall digestibility without negative impact on growth and development. Thus, grass biomass utilization may potentially be enhanced by manipulation of the flavone biosynthesis pathway.







Leave a Reply
Want to join the discussion?Feel free to contribute!