The Terpene Synthases of Red Algae Have a Bacterial Origin
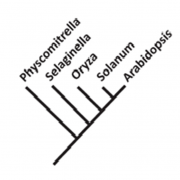
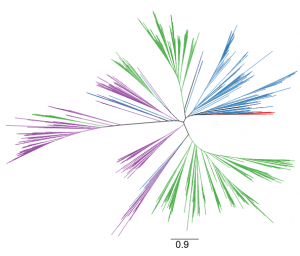 The red algae (Rhodophyta), encompassing over 8,000 species, are the richest source of marine secondary metabolites. Among red algae, many genera produce terpenes, which constitute the largest class of secondary metabolites. Despite the rich diversity of terpenes in red algae, little is known about how they are biosynthesized. This is in sharp contrast to our considerable knowledge of terpene biosynthesis in land plants. Land plants produce a large array of terpenes, whose skeletal diversity can be attributed largely to terpene synthases. Two types of terpene synthases are known: typical plant terpene synthases and microbial terpene synthase-like (MTPSL) enzymes. The genes encoding typical plant terpene synthase are well understood. In contrast, MTPSL genes were discovered only recently and are more closely related to terpene synthase genes from bacteria and fungi than to typical plant terpene synthase genes. By systematic sequence analysis of seven genomes and 34 transcriptomes of red algae, Wei et al. (10.1104/pp.18.01413) have identified MTPSL homologs within one genome and two transcriptomes of red algae: no homologs of typical plant terpene synthase genes were found. Phylogenetic analysis showed that red algae MTPSLs group with bacterial terpene synthases. Analysis of the genome assembly and characterization of neighboring genes demonstrated that red algal MTPSLs are bona fide red algal genes and not microbial contaminants. MTPSL genes from Porphyridium purpureum and Erythrolobus australicus were characterized via heterologous expression and demonstrated to have sesquiterpene synthase activities. Expression of the MTPSL gene in P. purpureum was found to be induced by methyl jasmonate, suggesting a role for this gene in host defense. In summary, this study indicates that the formation of terpene carbon skeletons in red algae is carried out by MTPSLs that are phylogenetically unrelated to typical plant terpene synthases and most likely originated in Rhodophyta via horizontal gene transfer from bacteria.
The red algae (Rhodophyta), encompassing over 8,000 species, are the richest source of marine secondary metabolites. Among red algae, many genera produce terpenes, which constitute the largest class of secondary metabolites. Despite the rich diversity of terpenes in red algae, little is known about how they are biosynthesized. This is in sharp contrast to our considerable knowledge of terpene biosynthesis in land plants. Land plants produce a large array of terpenes, whose skeletal diversity can be attributed largely to terpene synthases. Two types of terpene synthases are known: typical plant terpene synthases and microbial terpene synthase-like (MTPSL) enzymes. The genes encoding typical plant terpene synthase are well understood. In contrast, MTPSL genes were discovered only recently and are more closely related to terpene synthase genes from bacteria and fungi than to typical plant terpene synthase genes. By systematic sequence analysis of seven genomes and 34 transcriptomes of red algae, Wei et al. (10.1104/pp.18.01413) have identified MTPSL homologs within one genome and two transcriptomes of red algae: no homologs of typical plant terpene synthase genes were found. Phylogenetic analysis showed that red algae MTPSLs group with bacterial terpene synthases. Analysis of the genome assembly and characterization of neighboring genes demonstrated that red algal MTPSLs are bona fide red algal genes and not microbial contaminants. MTPSL genes from Porphyridium purpureum and Erythrolobus australicus were characterized via heterologous expression and demonstrated to have sesquiterpene synthase activities. Expression of the MTPSL gene in P. purpureum was found to be induced by methyl jasmonate, suggesting a role for this gene in host defense. In summary, this study indicates that the formation of terpene carbon skeletons in red algae is carried out by MTPSLs that are phylogenetically unrelated to typical plant terpene synthases and most likely originated in Rhodophyta via horizontal gene transfer from bacteria.