Advances in Understanding Root Hair Formation
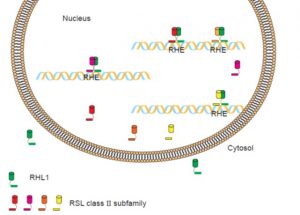 Root hairs greatly increase the surface area of roots, thereby facilitating the uptake of nutrients and water from the rhizosphere. They also serve as sites for plant interactions with soil microorganisms. Thus, elucidation of the molecular pathway for their development is important for potential modification of root hair morphology to produce crops with improved growth traits. ROOT HAIR DEFECTIVE SIX-LIKE (RSL) class II proteins are expressed preferentially in future root hair cells of Arabidopsis. Moon et al. (10.1104/pp.18.01002) functionally characterized the seven members of the RSL Class II subfamily in rice (Oryza sativa) genome: six of these genes were preferentially expressed and four were strongly expressed in root hairs. Overexpression (OX) of each of the four highly expressed family members in rice resulted in an increase in the density and length of root hairs. These four members also possess a transcription activation domain and are targeted to the nucleus. Genetic mutation of one member (Os07g39940) caused a severe reduction in root hair length. Biochemical studies demonstrated that the RSL Class II members interact with Root Hairless1 (OsRHL1), a key regulator of root hair development, and apparently assist in its translocation into the nucleus. A transcriptome analysis of Os07g39940-OX plants revealed that 86 genes, including Class III peroxidases, were highly up-regulated. Furthermore, reactive oxygen species levels in the root hairs were increased in Os07g39940-OX plants but were drastically reduced in the os07g39940 and rhl1 mutants. These results demonstrate that RSL Class II members are essential regulators of root hair development in rice.
Root hairs greatly increase the surface area of roots, thereby facilitating the uptake of nutrients and water from the rhizosphere. They also serve as sites for plant interactions with soil microorganisms. Thus, elucidation of the molecular pathway for their development is important for potential modification of root hair morphology to produce crops with improved growth traits. ROOT HAIR DEFECTIVE SIX-LIKE (RSL) class II proteins are expressed preferentially in future root hair cells of Arabidopsis. Moon et al. (10.1104/pp.18.01002) functionally characterized the seven members of the RSL Class II subfamily in rice (Oryza sativa) genome: six of these genes were preferentially expressed and four were strongly expressed in root hairs. Overexpression (OX) of each of the four highly expressed family members in rice resulted in an increase in the density and length of root hairs. These four members also possess a transcription activation domain and are targeted to the nucleus. Genetic mutation of one member (Os07g39940) caused a severe reduction in root hair length. Biochemical studies demonstrated that the RSL Class II members interact with Root Hairless1 (OsRHL1), a key regulator of root hair development, and apparently assist in its translocation into the nucleus. A transcriptome analysis of Os07g39940-OX plants revealed that 86 genes, including Class III peroxidases, were highly up-regulated. Furthermore, reactive oxygen species levels in the root hairs were increased in Os07g39940-OX plants but were drastically reduced in the os07g39940 and rhl1 mutants. These results demonstrate that RSL Class II members are essential regulators of root hair development in rice.