The Small Secreted Peptides of Legumes
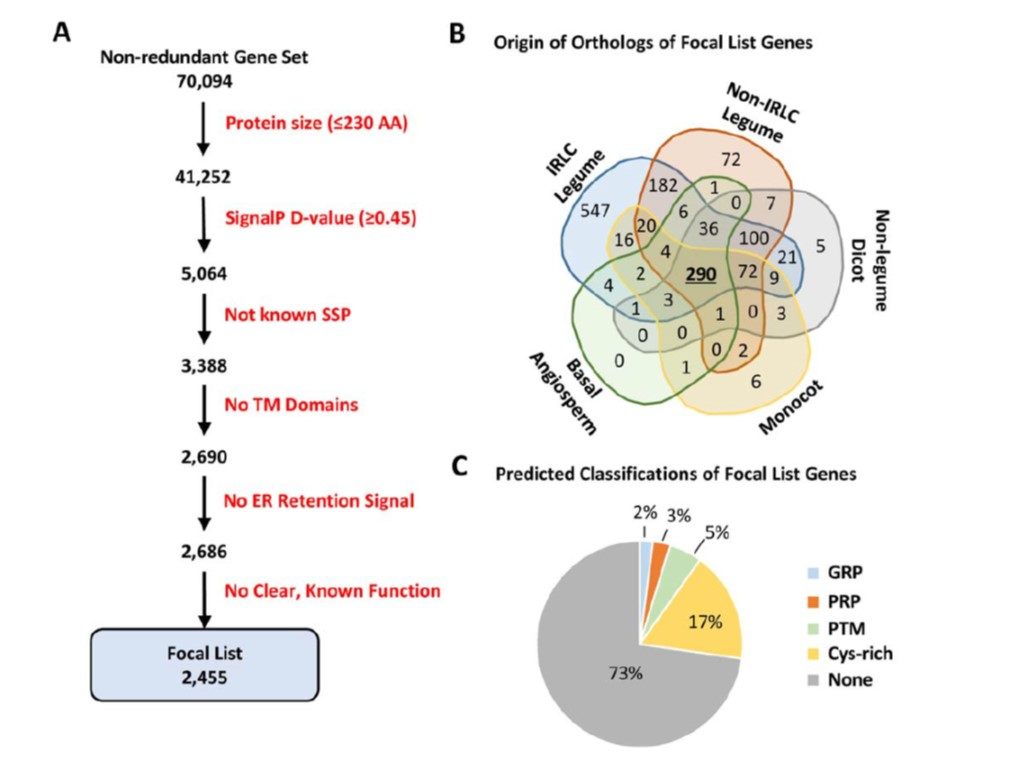
Small secreted peptides (SSPs) are peptides of 5 to 50 amino acid residues that are secreted into the apoplast. SSPs are critical regulators of a diverse array of growth and developmental processes in plants, including root growth, nutrient homeostasis, meristem maintenance, stress acclimation, pathogen defense, reproductive development, and symbiotic interactions. Despite the importance of SSPs, the vast majority remain unstudied and, indeed, unannotated in plant genomes. This is because gene prediction algorithms are biased against smaller gene products, to avoid wrongful annotation of random or noncoding transcripts, thereby excluding most SSPs and other bona fide small proteins. Moreover, many SSPs and other small proteins that are annotated have been identified merely as unknown proteins or hypothetical proteins, which, in practice, hinders identification of their function. Medicago truncatula has emerged as useful model for studies of legume biology and symbiosis, yet the SSP-coding potential of M. truncatula, like most other plant species, remains unclear. Moreover, only a tiny fraction of the known SSP-coding genes in M. truncatula have been connected to specific functions. deBang et al. (10.1104/pp.17.01096) have performed a systematic reannotation of the M. truncatula genome and succeeded in identifying 1,970 homologs of established SSP gene families and an additional 2,455 genes that are potentially novel SSPs. The authors were also able to identify hundreds of SSP genes regulated by macronutrient stress, rhizobial inoculation, and nodule development. Synthetic peptide applications were shown to alter root growth and nodulation phenotypes, revealing additional regulators of legume nutrient acquisition. These results should enable further investigations of specific SSP functions via peptide treatment and reverse genetics.