SUMOylation and the Regulation of Aluminum Resistance
Fang and Zhang et al. show that SUMOylation and deSUMOylation regulate the stability of a transcription factor, thereby influencing aluminum resistance in Arabidopsis. Descriptive paragraph. Plant Cell https://doi.org/10.1105/tpc.20.00687
Background: Aluminum (Al) is a primary constraint for crop production on acid soils, which make up more than 30% of the arable land in the world. Plants have evolved resistance mechanisms to detoxify Al. The model plant Arabidopsis thaliana utilizes the essential transcription factor STOP1 to regulate Al resistance. STOP1 protein has been reported to be regulated by ubiquitination, a type of post-translational modification.
Question: We wanted to know if STOP1 can be modified by other post-translational marks, and if so, we wanted to know how these modifications affect STOP1 protein function and Al resistance.
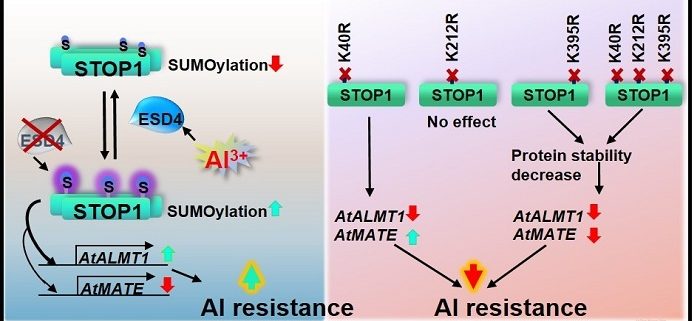
Findings: We found that STOP1 in Arabidopsis thaliana can be mono-SUMOylated at three lysine (K) sites: K40, K212 or K395, and deSUMOylated by the SUMO protease ESD4. When we mutated ESD4, the STOP1 SUMOylation level was higher and STOP1 protein function was changed, which caused the higher expression of the key STOP1-regulated gene AtALMT1 and ultimately increased Al resistance in the esd4 mutants. Conversely, when the STOP1 SUMOylation was prevented, the STOP1 protein level was lower and consequently AtALMT1 expression was decreased, which led to reduced Al resistance.
Next steps: How does Al stress increase STOP1 protein levels? We predict that other post-translational modifications on STOP1 are involved. Uncovering additional layers of post-translational regulation and dissecting the interaction among these potential regulatory mechanisms will be a stepping stone to engineer Al resistance in crops in the future.
Qiu Fang, Jie Zhang, Yang Zhang, Ni Fan, Harrold A. van den Burg, and Chao-Feng Huang. (2019). Regulation of Aluminum-Resistance in Arabidopsis Involves the SUMOylation of the Zinc Finger Transcription Factor STOP1. Plant Cell. https://doi.org/10.1105/tpc.20.00687