New Insights into Wound-Induced Callus Formation
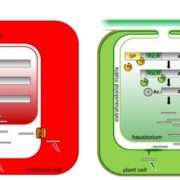
 Plants repair wound sites through the formation of unorganized cell masses called calli, which can also serve as progenitors of new organs. Callus formation and organ regeneration often entail cell cycle re-entry of quiescent cells, which is achieved through the re-activation of core cell cycle regulators CYCLIN (CYC) and CYCLIN-DEPENDENT KINASES (CDK). Recent studies have revealed how cells transduce wound signals to activate cell proliferation and callus formation. A set of AP2/ERF transcription factors called WOUND INDUCED DEDIFFERENTIATION1 (WIND1), WIND2, WIND3 and WIND4 plays key roles in wound-induced callus formation. Ikeuchi et al. (10.1104/pp.17.01035) have combined transcriptome analysis with quantitative hormonal analysis to investigate how wounding induces callus formation in Arabidopsis. A time-course analysis revealed that wounding induces dynamic transcriptional changes, starting from rapid stress responses followed by the activation of metabolic processes and protein synthesis and subsequent activation of cell cycle regulators. Gene ontology analyses suggest that wounding modifies the expression of hormone biosynthesis and response genes. A quantitative analysis of endogenous plant hormones revealed the accumulation of cytokinin prior to callus formation. Mutants defective in cytokinin synthesis or signaling display reduced efficiency in callus formation, indicating that de novo synthesis of cytokinin is critical for wound-induced callus formation. Other evidence presented suggests a possible role for ETHYLENE RESPONSE FACTOR 115 (ERF115) in callus formation. The authors demonstrate that type-B ARABIDOPSIS RESPONSE REGULATOR (ARR)-mediated cytokinin signaling regulates the expression of CYCLIN D3;1 (CYCD3;1) and that mutations in CYCD3;1 and its homologs CYCD3;2-3 cause defects in callus formation.. In toto, the results presented provide novel mechanistic insights into how wounding reactivates cell proliferation during callus formation.
Plants repair wound sites through the formation of unorganized cell masses called calli, which can also serve as progenitors of new organs. Callus formation and organ regeneration often entail cell cycle re-entry of quiescent cells, which is achieved through the re-activation of core cell cycle regulators CYCLIN (CYC) and CYCLIN-DEPENDENT KINASES (CDK). Recent studies have revealed how cells transduce wound signals to activate cell proliferation and callus formation. A set of AP2/ERF transcription factors called WOUND INDUCED DEDIFFERENTIATION1 (WIND1), WIND2, WIND3 and WIND4 plays key roles in wound-induced callus formation. Ikeuchi et al. (10.1104/pp.17.01035) have combined transcriptome analysis with quantitative hormonal analysis to investigate how wounding induces callus formation in Arabidopsis. A time-course analysis revealed that wounding induces dynamic transcriptional changes, starting from rapid stress responses followed by the activation of metabolic processes and protein synthesis and subsequent activation of cell cycle regulators. Gene ontology analyses suggest that wounding modifies the expression of hormone biosynthesis and response genes. A quantitative analysis of endogenous plant hormones revealed the accumulation of cytokinin prior to callus formation. Mutants defective in cytokinin synthesis or signaling display reduced efficiency in callus formation, indicating that de novo synthesis of cytokinin is critical for wound-induced callus formation. Other evidence presented suggests a possible role for ETHYLENE RESPONSE FACTOR 115 (ERF115) in callus formation. The authors demonstrate that type-B ARABIDOPSIS RESPONSE REGULATOR (ARR)-mediated cytokinin signaling regulates the expression of CYCLIN D3;1 (CYCD3;1) and that mutations in CYCD3;1 and its homologs CYCD3;2-3 cause defects in callus formation.. In toto, the results presented provide novel mechanistic insights into how wounding reactivates cell proliferation during callus formation.