Is All Root Hair Development the Same?
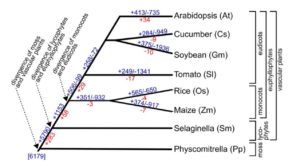 Root hairs are long tubular extensions of root epidermal cells that greatly increase the root surface area and thereby assist in water and nutrient absorption. Root hairs are found in nearly all vascular plants, including angiosperms, gymnosperms, and lycophytes, and they exhibit similar cellular features, suggesting a common evolutionary origin. However, different plant species are known to vary in their root hair distribution patterns and their root hair morphology, implying that genetic differences exist in root hair development programs. Root hairs have been studied intensively in Arabidopsis. In particular, molecular genetic analyses have led to the identification of numerous root hair genes, which provide insight into the mechanisms of Arabidopsis root hair development. Root hair-bearing cells in Arabidopsis are specified by a set of early-acting patterning genes that generate a cell position-dependent distribution of root hair cells and non-hair cells via a complex transcriptional regulatory network. To understand the extent to which this program might operate in other plants, Huang et al. (10.1104/pp.17.00374) conducted a large-scale comparative analysis of root hair development genes from diverse vascular plants, including eudicots, monocots, and a lycophyte. Combining phylogenetics and transcriptomics, the authors have discovered conservation of a core set of root hair genes across all vascular plants, which may derive from an ancient program for unidirectional cell growth coopted for root hair development during vascular plant evolution. Interestingly, they also discovered diversification in the structure and expression of root hair development genes, relative to other root hair- and root-expressed genes, among these species. The greatest divergence appears to have occurred in the composition and expression of genes used for root hair patterning, suggesting that the Arabidopsis transcriptional regulatory mechanism is not shared by other species. Altogether, this broad analysis of gene expression in a single cell type across multiple species provides new insight into the conservation and diversification of plant cell differentiation programs in vascular plants.
Root hairs are long tubular extensions of root epidermal cells that greatly increase the root surface area and thereby assist in water and nutrient absorption. Root hairs are found in nearly all vascular plants, including angiosperms, gymnosperms, and lycophytes, and they exhibit similar cellular features, suggesting a common evolutionary origin. However, different plant species are known to vary in their root hair distribution patterns and their root hair morphology, implying that genetic differences exist in root hair development programs. Root hairs have been studied intensively in Arabidopsis. In particular, molecular genetic analyses have led to the identification of numerous root hair genes, which provide insight into the mechanisms of Arabidopsis root hair development. Root hair-bearing cells in Arabidopsis are specified by a set of early-acting patterning genes that generate a cell position-dependent distribution of root hair cells and non-hair cells via a complex transcriptional regulatory network. To understand the extent to which this program might operate in other plants, Huang et al. (10.1104/pp.17.00374) conducted a large-scale comparative analysis of root hair development genes from diverse vascular plants, including eudicots, monocots, and a lycophyte. Combining phylogenetics and transcriptomics, the authors have discovered conservation of a core set of root hair genes across all vascular plants, which may derive from an ancient program for unidirectional cell growth coopted for root hair development during vascular plant evolution. Interestingly, they also discovered diversification in the structure and expression of root hair development genes, relative to other root hair- and root-expressed genes, among these species. The greatest divergence appears to have occurred in the composition and expression of genes used for root hair patterning, suggesting that the Arabidopsis transcriptional regulatory mechanism is not shared by other species. Altogether, this broad analysis of gene expression in a single cell type across multiple species provides new insight into the conservation and diversification of plant cell differentiation programs in vascular plants.



Leave a Reply
Want to join the discussion?Feel free to contribute!