Auxin Regulates Growth of a Characean Alga
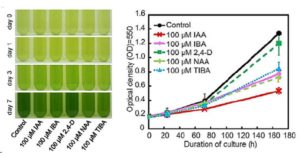 Auxin regulates many aspects of growth and development in land plants, but the origin and evolution of auxin signaling and response mechanisms remain largely unknown. Genome analyses of the moss Physcomitrella patens revealed the presence of the principal gene families involved in auxin homeostasis and signaling in tracheophytes, suggesting that the last common ancestor of land plants had already acquired the core auxin machinery of land plants. However, as we peer further back into phylogenetic time, things get murkier. To address this knowledge deficit, Ohtaka et al. () analyzed auxin responses in the charophyte alga Klebsormidium nitens, whose ancestor diverged from a green algal ancestor during the evolution of land plants. The authors have previously identified gene homologs for several auxin-biosynthesis and auxin signaling–related factors (TAA, YUCCA, PIN, AUX/LAX, and ABP1) in K. nitens; furthermore, the auxin indole-3-acetic acid (IAA) has been detected in K. nitens. On the other hand, a draft genome sequence suggested that K. nitens lacks the canonical TIR1-Aux/IAA-ARF–mediated auxin-signaling pathway found in land plants. The authors now report that exogenous IAA inhibits cell division and cell elongation in K. nitens as do inhibitors of auxin biosynthesis and inhibitors of polar auxin transport. Moreover, exogenous IAA rapidly induced expression of the LATERAL ORGAN BOUNDARIES-DOMAIN (LBD) transcription factor. These results suggest that K. nitens has acquired the part of the auxin system that regulates transcription and cell growth without the requirement for the central players that govern auxin signaling in land plants.
Auxin regulates many aspects of growth and development in land plants, but the origin and evolution of auxin signaling and response mechanisms remain largely unknown. Genome analyses of the moss Physcomitrella patens revealed the presence of the principal gene families involved in auxin homeostasis and signaling in tracheophytes, suggesting that the last common ancestor of land plants had already acquired the core auxin machinery of land plants. However, as we peer further back into phylogenetic time, things get murkier. To address this knowledge deficit, Ohtaka et al. () analyzed auxin responses in the charophyte alga Klebsormidium nitens, whose ancestor diverged from a green algal ancestor during the evolution of land plants. The authors have previously identified gene homologs for several auxin-biosynthesis and auxin signaling–related factors (TAA, YUCCA, PIN, AUX/LAX, and ABP1) in K. nitens; furthermore, the auxin indole-3-acetic acid (IAA) has been detected in K. nitens. On the other hand, a draft genome sequence suggested that K. nitens lacks the canonical TIR1-Aux/IAA-ARF–mediated auxin-signaling pathway found in land plants. The authors now report that exogenous IAA inhibits cell division and cell elongation in K. nitens as do inhibitors of auxin biosynthesis and inhibitors of polar auxin transport. Moreover, exogenous IAA rapidly induced expression of the LATERAL ORGAN BOUNDARIES-DOMAIN (LBD) transcription factor. These results suggest that K. nitens has acquired the part of the auxin system that regulates transcription and cell growth without the requirement for the central players that govern auxin signaling in land plants.





Leave a Reply
Want to join the discussion?Feel free to contribute!