Introducing the Plant Physiology Focus Issue on Cell Dynamics
Plant Physiology recently published a Focus Issue on Cell Dynamics. We asked the editors involved to tell us about what is meant by Cell Dynamics and why this topic is interesting and relevant, as well as about the Focus Issue program in general. This 11-minute video features Editor-in-Chief Mike Blatt (University of Glasgow), and Issue Editors Dan Szymanski (Purdue University), Diane Bassham (Iowa State University), and Wataru Sakamoto (Okayama University). Associate Features Editors Kim Johnson (La Trobe University), Emily Larson (University of Glasgow), and Mary Williams (ASPB) interviewed the editors.
The Focus Issue (January 2018) can be found here, and an editorial overview about it is here.
A transcript of the video can be found below.
Introduction by Editor in Chief Mike Blatt
[Mike] Hello, I’m Mike Blatt, I’m the Editor in Chief of Plant Physiology, and have been since 2013, and I’m here to talk to you very briefly about our most recent Focus Issue in Plant Cell Dynamics, and about our Focus Issue program in general.
Plant cell dynamics is a fascinating area of research, and it’s one that is attracting more and more attention in biology in general but certainly in all aspects of cellular function, plant function, and how plants interact with their environments.
It’s really driven by some amazing advances in technology that have taken place over the past two decades, and this now allows plant biologists, and biologists in general, to access the properties of cells and how they function in real time.
That has opened up huge areas of research that bridge the gaps between plant biology, biochemistry, biophysics and physical properties of cells, as well as the more traditional aspects of genetics and molecular biology.
So bringing all those together has really driven some exciting new advances that no doubt are going to change the way we think about how plants function and how they interact with their environment.
So I’m going to pass you over to the four editors who handled the Focus Issue, that’s Dan Szymanski, Diane Bassham, Wataru Sakamoto, and Teun Munnik. You will see they’re interviewed by several of our Assistant Features Editors, and they will be able to tell you a little bit about the focus issue on cell dynamics.
We run a Focus Issue program within Plant Physiology, and have been doing so for well over a decade now. Typically we’re publishing between two and four focus issues each year, with the idea of bringing together the most exciting and cutting-edge science and reviews of developments in the field along with primary research articles that go in that topic area. So these are great resources both for teaching and also for researchers who are interested in getting into an area and learning more about it first hand through experimentalists who are working in the field and can provide enough information to get researchers off the ground in starting their own studies.
And while obviously we’re focusing at the moment on Cell Dynamics, we have a number of other Focus Issues that are coming up in the near future. For example there will be one in this coming year on Biotic Interactions and we’re aiming for another Focus Issue on Synthetic Biology. You will find information about these already on our website, so if you are interested in submitting articles that would be relevant to either of those please look at the website or contact the editors to find out more about what they are looking for.
Thanks very much, and I’ll pass you over now to our fantastic Cell Dynamicists!
Kim Johnson interviews Dan Szymanski
[Kim] Dan, you’re a Co-editor of new special issue in Plant Physiology focusing on Cell Dynamics. . Tell me about yourself and why you’re involved.
[Dan]. I’m a cell biologist at Purdue University. I’ve been working on cell morphogenesis for twenty years.
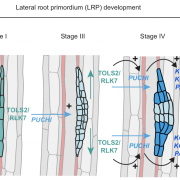
There was an opportunity to feature some of the hottest research in the area of cell biology so we put together a Focus Issue on the broad topic of cell dynamics. We’re really interested in emphasizing how the cell changed over time: different components and the Cellular and molecular mechanisms of shape change and trafficking, cell wall assembly.
Mary Williams interviews Wataru Sakamoto
[Mary] Could you tell us something about what is meant by Cell Dynamics, and your interest in this topic.
[Wataru] I am interested in organelles which are derived from endosymbiosis, namely mitochondria and chloroplasts. Those organelles contain more than two membrane systems, unlike the other organelles. And how these organelles develop is very fundamental, particularly for energy production. The chloroplast is very dynamic in the sense that it responds to light and moves, mitochondria as well, and this kind of dynamic nature is a very important part of cell dynamics.
Emily Larson interviews Diane Bassham
[Emily] With respect to the the topics that are being covered in this issue, how do you think about relating them to broader topics that might be interesting to non-scientists, or communicating it to non-scientists?
[Diane] Because cell biology is so fundamental to everything else that happens, and cells are really the defining feature of life. This means that we can relate cell biology and things that are happening in cells to almost any other aspect of biology including growth and development and even as far as ecological consideration. Think about stress responses and how the cell biology changes in different environmental conditions.
Mary and Wataru
[Mary] What are some of the more recent tools that have been developed that allow us to look at cells as dynamic structures?
[Wataru] Needless to say, most of the articles in this Focus Issue contain some part of organelle visualization. I think the most powerful tool that has been invented in the last two decades or so is live imaging analysis. In addition to that, as I mentioned the work by Yongliang Zhang, 3D construction is very important to look at the detailed structure of membranes.
Kim and Dan
[Kim] You’ve had all these advances, but what are some of the challenges that still remain to be addressed in cell biology and cell dynamics.
[Dan] I think part of it is getting the numbers out. For every hour you spend at the microscope collecting the images it can be anywhere from four to ten hours of analysing the images to get the quantitative data out of it.
I think one of the key challenges is image segmentation – extracting out quantitative information in an automated manner to increase the throughput of the experiment, because we’re talking about time-lapse imaging of the timescales of often hours. Plant cells are slow growing. So the datasets can grow quickly to gigabyte size. Dealing with that manually isn’t the most efficient solution.
The other thing is image registration. The cells move and tilt. How do you align the images so that you know where you’re at in the cell, so that over time you can properly reconstruct spatial coordinates over time. That’s a challenge.
Emily and Diane
[Emily] What is your opinion about where the most effort is needed right now?
[Diane] I think quantification is a going to be a huge issue. We have ways to quantify images, but there is a lot that still can be done when we’re thinking about fluorescence microscopy, even electron microscopy: the best ways to quantify, how to know what we’re quantifying, and what we’re really looking at. That’s essential for any kind of predictive work, you have to have accurate and extensive quantification.
Kim and Dan
[Kim] How is this knowledge going to help us address some of the big challenges that we’re facing today, food security and climate change. How will this help us maintain the productivity that we need?
[Dan] I think the future generations of crops are going to have to deal with unpredictable environments, and climate change, and new kinds of pathogens, viruses, bacterial and fungal pathogens.
There are major opportunities for engineering plants to have either new architectures or new pathways, to make them more adaptable to changing environments or more resistant to different types of pathogens.
These engineering approaches come at a price of understanding the underlying biology so all the work that’s been done and emphasized in this Focus Issue is trying to understand the key players in the plant cell that mediate these important traits.
So that’s really what it’s all about, is building this knowledge base from the ground up so that it can be used to understand traits in a more mechanistic way, to engineer traits rather than just select for them.
[Kim] Fantastic!