Review: The rice genome revolution: from an ancient grain to Green Super Rice ($) (Nat. Rev. Genetics)
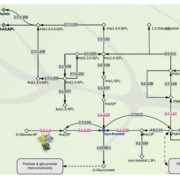
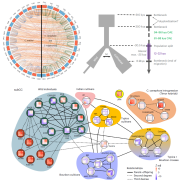
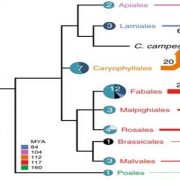
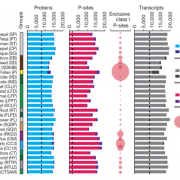
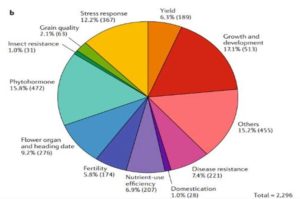 Rice is one of the major staple crops in the world as it is an essential component of diets and livelihoods. Populations in poor regions that are highly dependent on rice (Africa and South Asia) will increase dramatically by 2050, revealing the urgent need to find tools to prevent a future humanitarian crisis. Wing et al. reviewed how comparative and functional genomics of domesticated and wild rice germplasm collections provide valuable information for rice improvement. Genome mining can identify genes affecting growth and development, insect resistance, grain quality, stress response, yield, fertility, disease resistance and nutrient efficiency. The authors observe that various plant varieties have been crossed to obtain desirable phenotypes to increase the rice production. However, increasing production can also increase costs, as well as increase use of pesticides, fertilizers and water. Therefore, Green Super Rice (GSR) could be created combining characteristics of Asian rice (Oryza sativa) and African rice (Oryza glaberrima). If GSR could be produced, it could help to satisfy the demand for rice in resource-scarce regions, it might be economical in its production, and it could have the capacity to grow on marginal lands and with lower associated greenhouse gas emissions. (Summary by Gema Catalina Salas Leija and Maria Julissa Ek-Ramos). Nature Rev. Genetics 10.1038/s41576-018-0024-z.
Rice is one of the major staple crops in the world as it is an essential component of diets and livelihoods. Populations in poor regions that are highly dependent on rice (Africa and South Asia) will increase dramatically by 2050, revealing the urgent need to find tools to prevent a future humanitarian crisis. Wing et al. reviewed how comparative and functional genomics of domesticated and wild rice germplasm collections provide valuable information for rice improvement. Genome mining can identify genes affecting growth and development, insect resistance, grain quality, stress response, yield, fertility, disease resistance and nutrient efficiency. The authors observe that various plant varieties have been crossed to obtain desirable phenotypes to increase the rice production. However, increasing production can also increase costs, as well as increase use of pesticides, fertilizers and water. Therefore, Green Super Rice (GSR) could be created combining characteristics of Asian rice (Oryza sativa) and African rice (Oryza glaberrima). If GSR could be produced, it could help to satisfy the demand for rice in resource-scarce regions, it might be economical in its production, and it could have the capacity to grow on marginal lands and with lower associated greenhouse gas emissions. (Summary by Gema Catalina Salas Leija and Maria Julissa Ek-Ramos). Nature Rev. Genetics 10.1038/s41576-018-0024-z.