Epigenetic Divergence Associated with Heterosis
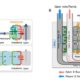
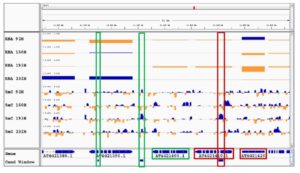 Heterosis refers to the tendency of a crossbred individual to show qualities superior to those of both parents. The phenomenon has been exploited extensively in agricultural breeding for decades and has improved crop performance enormously. Despite its commercial impact, knowledge of the molecular basis underlying heterosis remains incomplete. Most studies have focused on finding genetic explanations, resulting in the classical dominance and overdominance models of heterosis. Genetic explanations, however, do not sufficiently explain or predict heterosis. There is growing evidence that epigenetic factors also play a role in heterosis. Recent studies have implicated changes in DNA methylation and small RNAs in hybrid performance; however, it remains unclear whether epigenetic changes are a cause or a consequence of heterosis. Lauss et al. (pp. ***-***) have analyzed a large panel of over 500 Arabidopsis (Arabidopsis thaliana) epigenetic hybrid plants (epiHybrids), which they derived from near-isogenic but epigenetically divergent parents. This experimental system allowed them to quantify the contribution of parental methylation differences to heterosis. They measured traits such as leaf area, growth rate, flowering time, main stem branching, rosette branching, and final plant height and observed several strong positive and negative heterotic phenotypes among the epiHybrids. Using an epigenetic quantitative trait locus mapping approach, they succeeded in identifying specific differentially methylated regions in the parental genomes that are associated with hybrid performance. Sequencing of methylomes, transcriptomes, and genomes of selected parent-epiHybrid combinations further showed that these parental differentially methylated regions most likely mediate the remodeling of methylation and transcriptional states at specific loci in the hybrids. Taken together, their data suggest that locus-specific epigenetic divergence between the parental lines can directly or indirectly trigger heterosis in Arabidopsis hybrids independent of genetic changes. These results add to a growing body of evidence that points to epigenetic factors as one of the key determinants of hybrid performance.
Heterosis refers to the tendency of a crossbred individual to show qualities superior to those of both parents. The phenomenon has been exploited extensively in agricultural breeding for decades and has improved crop performance enormously. Despite its commercial impact, knowledge of the molecular basis underlying heterosis remains incomplete. Most studies have focused on finding genetic explanations, resulting in the classical dominance and overdominance models of heterosis. Genetic explanations, however, do not sufficiently explain or predict heterosis. There is growing evidence that epigenetic factors also play a role in heterosis. Recent studies have implicated changes in DNA methylation and small RNAs in hybrid performance; however, it remains unclear whether epigenetic changes are a cause or a consequence of heterosis. Lauss et al. (pp. ***-***) have analyzed a large panel of over 500 Arabidopsis (Arabidopsis thaliana) epigenetic hybrid plants (epiHybrids), which they derived from near-isogenic but epigenetically divergent parents. This experimental system allowed them to quantify the contribution of parental methylation differences to heterosis. They measured traits such as leaf area, growth rate, flowering time, main stem branching, rosette branching, and final plant height and observed several strong positive and negative heterotic phenotypes among the epiHybrids. Using an epigenetic quantitative trait locus mapping approach, they succeeded in identifying specific differentially methylated regions in the parental genomes that are associated with hybrid performance. Sequencing of methylomes, transcriptomes, and genomes of selected parent-epiHybrid combinations further showed that these parental differentially methylated regions most likely mediate the remodeling of methylation and transcriptional states at specific loci in the hybrids. Taken together, their data suggest that locus-specific epigenetic divergence between the parental lines can directly or indirectly trigger heterosis in Arabidopsis hybrids independent of genetic changes. These results add to a growing body of evidence that points to epigenetic factors as one of the key determinants of hybrid performance.