Review: The genomic basis of adaptation in plants ($)
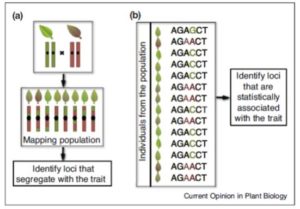 Evolution starts with molecular variation and phenotypic diversity, upon which selection acts. Flood and Hancock review the approaches used to detect adaptive evolution. The top down approach starts with the phenotype and works to identify its genomic basis; examples are quantitative trait locus (QTL) mapping or genome-wide association screening (GWAS). The bottom up approach starts with genomic polymorphisms and connects them to adaptive phenotypes, through population genomics and genomic scan approaches. Patterns of polymorphisms and linkage disequilibria can shed light on the nature of the adaptive scenario, for example whether selection involved a hard or soft sweep. As the authors describe, these approaches and models can be applied to both natural selection and adaptation during domestication. Curr. Opin. Plant Biol. 10.1016/j.pbi.2017.02.003
Evolution starts with molecular variation and phenotypic diversity, upon which selection acts. Flood and Hancock review the approaches used to detect adaptive evolution. The top down approach starts with the phenotype and works to identify its genomic basis; examples are quantitative trait locus (QTL) mapping or genome-wide association screening (GWAS). The bottom up approach starts with genomic polymorphisms and connects them to adaptive phenotypes, through population genomics and genomic scan approaches. Patterns of polymorphisms and linkage disequilibria can shed light on the nature of the adaptive scenario, for example whether selection involved a hard or soft sweep. As the authors describe, these approaches and models can be applied to both natural selection and adaptation during domestication. Curr. Opin. Plant Biol. 10.1016/j.pbi.2017.02.003





Leave a Reply
Want to join the discussion?Feel free to contribute!