An acidophilic green algal genome provides insights into adaptation to an acidic environment ($)
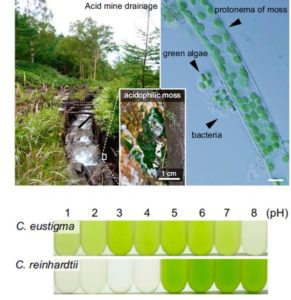 Hirooka et al. examined the genome of an acid-loving green alga, Chlamydomonas eustigma, to learn how it tolerates its low pH environment. Key differences between the acidophilic species and the neutrophilic species Chlamydomonas reinhardtii include: an increase in expression of genes encoding plasma membrane H+-ATPases (to better extrude protons) and heat-shock proteins (for molecular stress tolerance). Further, the acidophilic species has lost genes involved in organic acid-producing fermentation pathways, and has acquired genes involved in arsenic detoxification through horizontal gene transfer; arsenic is more readily taken up in acidic environments. This study extends our understanding of the requirements for life at low pH. The authors also suggest that acidophilic algae can be useful for food or fuel cultivation outdoors, as they can be grown in acidic conditions without risk of contamination by other organisms. Proc. Natl. Acad. Sci. USA 10.1073/pnas.1707072114.
Hirooka et al. examined the genome of an acid-loving green alga, Chlamydomonas eustigma, to learn how it tolerates its low pH environment. Key differences between the acidophilic species and the neutrophilic species Chlamydomonas reinhardtii include: an increase in expression of genes encoding plasma membrane H+-ATPases (to better extrude protons) and heat-shock proteins (for molecular stress tolerance). Further, the acidophilic species has lost genes involved in organic acid-producing fermentation pathways, and has acquired genes involved in arsenic detoxification through horizontal gene transfer; arsenic is more readily taken up in acidic environments. This study extends our understanding of the requirements for life at low pH. The authors also suggest that acidophilic algae can be useful for food or fuel cultivation outdoors, as they can be grown in acidic conditions without risk of contamination by other organisms. Proc. Natl. Acad. Sci. USA 10.1073/pnas.1707072114.