Understanding leaf shape development and diversity in Brassicaceae (New Phytol) ($)
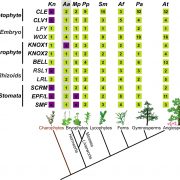
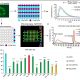
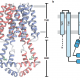
 Many economically important crops are in the Brassicaceae family, such as cabbage, mustard, and the model plant Arabidopsis thaliana. Recently, the systematics of Brassicaceae has assigned most of the species to 52 monophyletic groupings (tribes). However, relationships along the backbone of the phylogeny and among tribes remain largely unknown because of the lack of resolution. To break this limitation, Nikolov et al. reconstructed a robust phylogenetic framework together with the comparative transcriptome analysis to study the leaf shape evolution in the Brassicaceae family. Their phylogenomic study benefits the future genomic, developmental, and evolutionary studies among the Brassicaceae family, which allows us to understand the genetic basis for the evolution of development in crucifers. (Summary and grapical abstract by Jin Liao) New Phytol. 10.1111/nph.15732
Many economically important crops are in the Brassicaceae family, such as cabbage, mustard, and the model plant Arabidopsis thaliana. Recently, the systematics of Brassicaceae has assigned most of the species to 52 monophyletic groupings (tribes). However, relationships along the backbone of the phylogeny and among tribes remain largely unknown because of the lack of resolution. To break this limitation, Nikolov et al. reconstructed a robust phylogenetic framework together with the comparative transcriptome analysis to study the leaf shape evolution in the Brassicaceae family. Their phylogenomic study benefits the future genomic, developmental, and evolutionary studies among the Brassicaceae family, which allows us to understand the genetic basis for the evolution of development in crucifers. (Summary and grapical abstract by Jin Liao) New Phytol. 10.1111/nph.15732