Plants with self-sustained luminescence (bioRxiv)
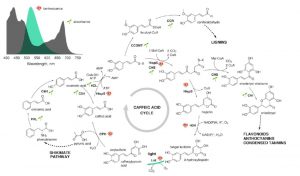 Nothing beats being able see gene expression in real time and space. In recent years, plant biologists have made great strides in understanding plants by using the visual reporters GUS, green fluorescent protein (and other fluorescent proteins) and luciferase. Each of these requires either a substrate or an illuminating light source. Now, Mitiouchkina et al. have developed plants that luminesce without requiring any exogenous inputs. They started with a luminescent pathway from fungi that draws substrates from the plant’s own caffeic acid pathway. Here they present our first views of this new method, with an overview of where highest levels of luminescence are observed in the plant, and insights into the rate-limiting substrate. It will interesting to see how quickly this new reporter is embraced and optimized by the community! (Summary by Mary Williams) bioRxiv 10.1101/809376.
Nothing beats being able see gene expression in real time and space. In recent years, plant biologists have made great strides in understanding plants by using the visual reporters GUS, green fluorescent protein (and other fluorescent proteins) and luciferase. Each of these requires either a substrate or an illuminating light source. Now, Mitiouchkina et al. have developed plants that luminesce without requiring any exogenous inputs. They started with a luminescent pathway from fungi that draws substrates from the plant’s own caffeic acid pathway. Here they present our first views of this new method, with an overview of where highest levels of luminescence are observed in the plant, and insights into the rate-limiting substrate. It will interesting to see how quickly this new reporter is embraced and optimized by the community! (Summary by Mary Williams) bioRxiv 10.1101/809376.