Keeping a lid on shoot regeneration: SIZ1 suppresses wound-induced developmental reprogramming
Michael J. Skelly
ORCID ID: 0000-0002-9024-0037
Institute of Molecular Plant Sciences, School of Biological Sciences, University of Edinburgh, Edinburgh EH9 3BF, United Kingdom
Plants have the remarkable ability to regenerate tissues and organs in response to wounding. This process relies on the formation of callus, a mass of unorganised cells that provides protection to the wounding site while allowing cell reprogramming and proliferation. Callus can be formed in vitro by incubating explants on auxin-rich callus-inducing medium (CIM), which stimulates reactivation of lateral root formation (Sugimoto et al., 2010). Subsequently, callus can be transferred to cytokinin-rich shoot-inducing medium (SIM), which triggers reprogramming of callus cells to shoot meristem identity and allows regeneration of a whole plant. Extensive transcriptional networks involving a wide range of transcription factors, hormones and additional regulators have been identified that control each step of these developmental transitions. However, it has remained unknown if post-translational regulation occurs during these wounding and developmental processes.
Various post-translational protein modifications (PTMs) increase the size and functionality of proteomes, enabling exquisite control of stress responses and signalling pathways. SUMOylation, the conjugation of the SMALL UBIQUITIN-LIKE MODIFIER (SUMO) to target proteins, is a well-characterised PTM that controls the activity of many important transcriptional regulators in plants (Augustine and Vierstra, 2018). SUMO is conjugated to proteins by an enzymatic cascade involving a SUMO-activating E1 enzyme, a SUMO-conjugating E2 enzyme, and SUMO E3 ligases. The Arabidopsis thaliana genome encodes two SUMO E3 ligases: METHYL METHANESULFONATE-SENSITIVE 21 (MMS21) and SAP AND MIZ1 (SIZ1). Of these, SIZ1 is the predominant E3 ligase responsible for the majority of SUMO ligation to target proteins (Rytz et al., 2018). While SUMOylation has been implicated in some developmental processes, it is unknown whether it functions in wounding-induced regeneration.
In this issue of Plant Physiology, Coleman et al. (2020) report that SIZ1 supresses in vitro shoot regeneration in Arabidopsis. The authors compared the shoot regeneration capacities of wild-type and siz1 mutant plants by incubating explants on CIM before transferring them to SIM. Shoot regeneration was dramatically enhanced in siz1 mutants, with new shoots appearing earlier and in greater numbers than in wild type. Expression of a SIZ1 transgene under its native promoter rescued this enhanced shoot generation phenotype of siz1 plants. At the time of transfer between CIM and SIM, callus size and morphology were indistinguishable between wild type and siz1 mutants. These results suggest that SIZ1 negatively regulates in vitro shoot regeneration.
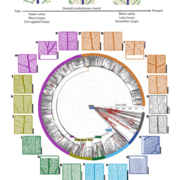
To uncover any transcriptomic differences between wild type and siz1 mutants, the authors performed RNA-sequencing (RNA-seq) at different time points during shoot regeneration. The difference in global gene expression was most apparent immediately after cutting, with 1,375 genes upregulated and 912 genes downregulated in siz1 compared to wild type. After 4 days of incubation on CIM, the difference between genotypes was much less pronounced, but after transferring the cuttings to SIM, siz1 plants again showed profound differences in gene expression compared to wild type at both 4 and 6 days post-transfer. The mis-expressed genes in siz1 plants were mostly distinct after either cutting, incubation on CIM or SIM. Gene ontology (GO) analyses revealed that genes upregulated in siz1 after cutting were largely associated with stress responses. The authors compared these genes to those identified as upregulated in intact siz1 plants in other studies (Catala et al., 2007; Rytz et al., 2018) and established that most of the upregulated genes in the present study were unique to cutting. Together these data suggest that siz1 mutants display a heightened stress response to wounding. Accordingly, in the absence of any external hormone treatment, siz1 mutants displayed enhanced callus formation compared to wild-type plants after cutting of hypocotyls.
Wounding responses in plants are regulated by a range of phytohormones including salicylic acid (SA), jasmonic acid, abscisic acid, and ethylene. Many genes representing GO categories related to all of these hormones were upregulated in siz1 mutants after cutting. It was previously shown that the dwarf phenotypes and autoimmunity of siz1 mutants were due to high levels of SA and these phenotypes were reversible by expression of a bacterial salicylate hydrolase, NahG, which degrades SA (Lee et al., 2007). To determine if the enhanced shoot regeneration phenotype of siz1 is a result of high SA levels, Coleman et al. (2020) compared siz1 plants expressing NahG to siz1 single mutants. No difference in shoot regeneration was observed between these genotypes, suggesting that the enhanced shoot regeneration of siz1 mutants is independent of SA signalling.
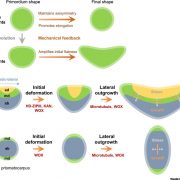
The ability of plant tissues to form callus and acquire regeneration capacity after wounding is promoted by the transcription factor WOUND-INDUCED DEDIFFERENTIATION 1 (WIND1) (Iwase et al., 2011). Coleman et al. (2020) report that expression of both WIND1 and its homologue WIND2 are elevated in siz1 mutants after cutting. To determine if the enhanced shoot regeneration observed in siz1 mutants was due to elevated WIND1 expression, the authors expressed a dominant-negative form of WIND1 fused to a SUPERMAN repression domain (SRDX) in siz1 plants. These siz1-3 WIND1-SRDX plants showed less shoot regeneration than siz1-3 plants but this phenotype was not rescued to wild-type levels. This suggests that the enhanced shoot regeneration capacity of siz1 plants is partially due to enhanced WIND1 activity. The authors further investigated this by comparing genes upregulated in 35S::WIND1 seedlings from a previous study (Iwase et al., 2011) to those upregulated in siz1 at 4 days on CIM. There was a substantial overlap of genes, further suggesting that the enhanced shoot regeneration of siz1 plants is due to a hyperactive wounding response.
The authors also examined transcriptional responses to hormones that are involved in both the acquisition of regeneration capacity and in vitro shoot regeneration. While overall transcriptional responses to auxin were unaffected in siz1, transcription of cytokinin-responsive genes was altered in siz1 plants. This altered transcription included enhanced expression of the key regulator of shoot meristem induction WUSCHEL (WUS). Compared to wild type, siz1 mutants were hypersensitive to exogenously applied cytokinin. Together, these data suggest that SIZ1 is involved in regulating the cytokinin response in the context of shoot meristem regeneration.
In summary, this exciting study by Coleman et al. (2020) shows that in vitro shoot regeneration is suppressed by SIZ1-mediated regulation of the transcriptional wounding response. While loss of SIZ1 function does not appear to affect auxin-induced callus formation or pluripotency, it enhances cytokinin-induced shoot meristem regulators. It is unclear how SIZ acts on this pathway and further work should focus on identifying SUMO substrates modified by SIZ1 during the different stages of in vitro shoot regeneration. This study did not examine any effects of the loss of SIZ1 function on global SUMOylation during shoot regeneration or identify any candidate proteins that might be SUMO substrates during this process. These experiments will be necessary in the future to understand how SIZ1 directly influences in vitro shoot regeneration and may uncover additional targets of SUMOylation that can be modulated to engineer shoot regeneration potential and/or stress tolerance in plants.
LITERATURE CITED
Augustine RC, Vierstra RD (2018) SUMOylation: re-wiring the plant nucleus during stress and development. Curr Opin Plant Biol 45: 143-154
Catala R, Ouyang J, Abreu IA, Hu Y, Seo H, Zhang X, Chua NH (2007) The Arabidopsis E3 SUMO ligase SIZ1 regulates plant growth and drought responses. Plant Cell 19: 2952-2966
Iwase A, Mitsuda N, Koyama T, Hiratsu K, Kojima M, Arai T, Inoue Y, Seki M, Sakakibara H, Sugimoto K, Ohme-Takagi M (2011) The AP2/ERF transcription factor WIND1 controls cell dedifferentiation in Arabidopsis. Curr Biol 21: 508-514
Lee J, Nam J, Park HC, Na G, Miura K, Jin JB, Yoo CY, Baek D, Kim DH, Jeong JC, Kim D, Lee SY, Salt DE, Mengiste T, Gong Q, Ma S, Bohnert HJ, Kwak SS, Bressan RA, Hasegawa PM, Yun DJ (2007) Salicylic acid-mediated innate immunity in Arabidopsis is regulated by SIZ1 SUMO E3 ligase. Plant J 49: 79-90
Rytz TC, Miller MJ, McLoughlin F, Augustine RC, Marshall RS, Juan YT, Charng YY, Scalf M, Smith LM, Vierstra RD (2018) SUMOylome Profiling Reveals a Diverse Array of Nuclear Targets Modified by the SUMO Ligase SIZ1 during Heat Stress. Plant Cell 30: 1077-1099
Sugimoto K, Jiao Y, Meyerowitz EM (2010) Arabidopsis regeneration from multiple tissues occurs via a root development pathway. Dev Cell 18: 463-471