Is Root Cortical Senescence Beneficial?
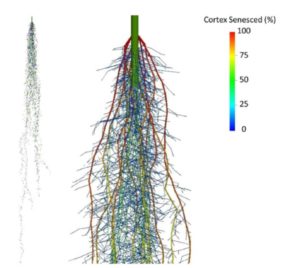 Root cortical senescence (RCS) is a type of programmed cell death found in the Triticeae tribe. RCS is unrelated to the formation of root cortical aerenchyma or the loss of the root cortex due to secondary growth in dicots. Conceivably RCS may benefit the plant by reducing maintenance respiration in the root or by nutrient reallocation from the senescing tissues. On the other hand, as RCS formation progresses and more cortical cells senescence, the continuity of the cell-to-cell pathway is increasingly disrupted. Additionally, RCS formation and root aging coincide with increased suberization of the endodermis which may also contribute to reduced hydraulic conductivity. To address the question of if and how RCS benefits plants, Schneider et al. () used the functional-structural model SimRoot to evaluate the functional implications of RCS in barley (Hordeum vulgare) under suboptimal nitrate, phosphorus, and potassium availability. By the time flowering occurred, RCS was projected to increase simulated plant growth by up to 52%, 73%, and 41% in nitrate, phosphorus, and potassium limiting conditions, respectively. Nutrient reallocation during RCS had a greater effect on simulated plant growth than reduced respiration or nutrient uptake. Additionally, RCS was quantified in field-grown barley in different nitrogen regimes. Living cortical volume per root length (an indicator of RCS) decreased with depth in younger plants, while roots of older plants had very little living cortical volume per root length. The authors conclude that RCS may be an adaptive trait for nutrient acquisition by reallocating nutrients from senescing tissue and secondarily by reducing root respiration. These simulated results suggest that RCS merits investigation as a breeding target for enhanced soil resource acquisition and edaphic stress tolerance in plants.
Root cortical senescence (RCS) is a type of programmed cell death found in the Triticeae tribe. RCS is unrelated to the formation of root cortical aerenchyma or the loss of the root cortex due to secondary growth in dicots. Conceivably RCS may benefit the plant by reducing maintenance respiration in the root or by nutrient reallocation from the senescing tissues. On the other hand, as RCS formation progresses and more cortical cells senescence, the continuity of the cell-to-cell pathway is increasingly disrupted. Additionally, RCS formation and root aging coincide with increased suberization of the endodermis which may also contribute to reduced hydraulic conductivity. To address the question of if and how RCS benefits plants, Schneider et al. () used the functional-structural model SimRoot to evaluate the functional implications of RCS in barley (Hordeum vulgare) under suboptimal nitrate, phosphorus, and potassium availability. By the time flowering occurred, RCS was projected to increase simulated plant growth by up to 52%, 73%, and 41% in nitrate, phosphorus, and potassium limiting conditions, respectively. Nutrient reallocation during RCS had a greater effect on simulated plant growth than reduced respiration or nutrient uptake. Additionally, RCS was quantified in field-grown barley in different nitrogen regimes. Living cortical volume per root length (an indicator of RCS) decreased with depth in younger plants, while roots of older plants had very little living cortical volume per root length. The authors conclude that RCS may be an adaptive trait for nutrient acquisition by reallocating nutrients from senescing tissue and secondarily by reducing root respiration. These simulated results suggest that RCS merits investigation as a breeding target for enhanced soil resource acquisition and edaphic stress tolerance in plants.






Leave a Reply
Want to join the discussion?Feel free to contribute!