Goldilocks Principle: MtNFH1 Ensures Optimal Nod Factor Activity
Partner selection is a critical step that must occur early during establishment of Root Nodule Symbiosis (RNS). RNS refers to the mutualistic interaction between legumes and some non-legumes with soil bacteria that help convert atmospheric nitrogen into plant usable ammonia. In legumes such as Medicago truncatula, a successful interaction initiates two genetically separable but interdependent developmental programs – Infection thread (IT) formation and nodule organogenesis. Only once the IT reaches the nodule cortex and the bacteria are released does nitrogen fixation commence (Oldroyd et al., 2011). However, for these processes to occur, legumes must first carefully choose their partner by exchanging chemical signals.
Plants secrete chemoattractants called flavonoids which in turn stimulate production of rhizobial strain-specific signaling molecules called Nod (Nodulation) factors (NFs). NFs comprise a chitin-based backbone of variable length that is acylated at the N-terminal end and can have additional functional group modifications resulting in unique chemical signatures (Wang et al., 2012). If rhizobia fail to produce the correct NF or produce too little, RNS is impaired (Roche et al., 1996). However, overproduction of NFs can also be detrimental to symbiosis. Cai et al. (2018) characterize the Medicago truncatula NOD FACTOR HYDROLASE 1 to demonstrate that not only the presence of specific NFs but also their concentration controls the establishment of RNS.
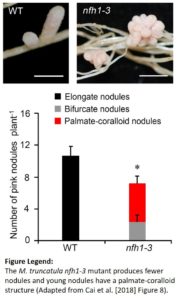 The Medicago truncatula NOD FACTOR HYDROLASE (MtNFH1) enzyme can hydrolyze the Sinorhizobium meliloti (Sm) NFs into lipo-disaccharides in vitro and in the rhizosphere (Tian et al. 2013). In the current study, the authors show that both the activity and the expression of MtNFH1 increase with increasing concentrations of the NFs but are strongly reduced in the dmi1, dmi2, and dmi3 mutants; indicating that MtNFH1 lies downstream of the common symbiosis pathway. Consistent with its localization in infection chambers of curled root hairs, mutants in this chitinase-like hydrolase gene had fewer infection events and the root hairs showed aberrant branching. The strongest mutant allele nfh1-3 had fewer nitrogen-fixing pink nodules and these nodules clustered together and possessed multiple meristems. The authors hypothesized that, since accumulation of uncleaved NFs resulting from a lack of NF hydrolase activity caused aberrant organogenesis, overproduction of NFs must similarly affect nodule morphology. In line with this reasoning, an Sm strain carrying extra copies of NF synthesis genes induced coralloid nodules with apical meristem branching on wild type roots.
The Medicago truncatula NOD FACTOR HYDROLASE (MtNFH1) enzyme can hydrolyze the Sinorhizobium meliloti (Sm) NFs into lipo-disaccharides in vitro and in the rhizosphere (Tian et al. 2013). In the current study, the authors show that both the activity and the expression of MtNFH1 increase with increasing concentrations of the NFs but are strongly reduced in the dmi1, dmi2, and dmi3 mutants; indicating that MtNFH1 lies downstream of the common symbiosis pathway. Consistent with its localization in infection chambers of curled root hairs, mutants in this chitinase-like hydrolase gene had fewer infection events and the root hairs showed aberrant branching. The strongest mutant allele nfh1-3 had fewer nitrogen-fixing pink nodules and these nodules clustered together and possessed multiple meristems. The authors hypothesized that, since accumulation of uncleaved NFs resulting from a lack of NF hydrolase activity caused aberrant organogenesis, overproduction of NFs must similarly affect nodule morphology. In line with this reasoning, an Sm strain carrying extra copies of NF synthesis genes induced coralloid nodules with apical meristem branching on wild type roots.
Curiously, transgenic overexpression of MtNFH1 yielded a similar phenotype as the mutant. These lines had fewer pink nodules and the ones that formed were branched. How can too much and too little of symbiotic NF be detrimental to establishment of RNS? The authors propose that MtNFH1 spatiotemporally limits NF levels in order to reach an optimal NF activity for initiating the right number of infections and correct nodule morphology. It remains to be seen how MtNFH1 relates to the phenomenon of autoregulation of nodulation in legumes, by which the plant controls the number of nodules formed.
REFERENCES
Cai, J., et al. (2018). Role of the Nod Factor Hydrolase MtNFH1 in Regulating Nod Factor Levels during Rhizobial Infection and in Mature Nodules of Medicago truncatula. Plant Cell. Published January 2018. DOI: https://doi.org/10.1105/tpc.17.00420.
Oldroyd, G.E.D., Murray, J.D., Poole, P.S., and Downie, J.A. (2011). The rules of engagement in the legume-rhizobial symbiosis. Annu Rev Genet 45, 119-144.
Roche, P., Maillet, F., Plazanet, C., Debelle, F., Ferro, M., Truchet, G., Prome, J.C., and Denarie, J. (1996). The common nodABC genes of Rhizobium meliloti are host-range determinants. Proc Natl Acad Sci U S A 93, 15305-15310.
Tian, Y., Liu, W., Cai, J., Zhang, L.Y., Wong, K.B., Feddermann, N., Boller, T., Xie, Z.P., and Staehelin, C. (2013). The nodulation factor hydrolase of Medicago truncatula: characterization of an enzyme specifically cleaving rhizobial nodulation signals. Plant Physiol 163, 1179–1190.
Wang, D., Yang, S.M., Tang, F., and Zhu, H.Y. (2012). Symbiosis specificity in the legume – rhizobial mutualism. Cell Microbiol 14, 334-342.