Global transcriptome analysis reveals circadian control of splicing events in Arabidopsis thaliana (bioRxiv)
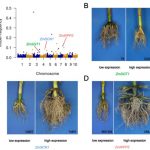
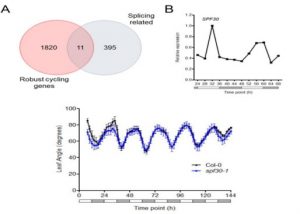 Developmental programming, metabolism and plant growth are regulated by the circadian rhythm that integrates environmental signals and plant growth. In this paper, Romanowski et al. have used the RNAseq approach to determine post-transcriptional regulation in Arabidopsis thaliana, specifically alternate splicing regulated by circadian rhythm. Tissues from 15-day old plants with altered light conditions (affecting growth), were harvested for deep sequencing. Using an in-house bioinformatics pipeline, the alternative splicing events were determined. In addition to the identification of novel regulated genes, the authors identified multiple transcripts undergoing post-transcriptional regulation. The authors also identified that the circadian rhythm regulates the spliceosome machinery in Arabidopsis by affecting the genes including SPLICING FACTOR 30 (AtSPF30). (Summary by Suresh Damodaran) bioRxiv 10.1101/845560.
Developmental programming, metabolism and plant growth are regulated by the circadian rhythm that integrates environmental signals and plant growth. In this paper, Romanowski et al. have used the RNAseq approach to determine post-transcriptional regulation in Arabidopsis thaliana, specifically alternate splicing regulated by circadian rhythm. Tissues from 15-day old plants with altered light conditions (affecting growth), were harvested for deep sequencing. Using an in-house bioinformatics pipeline, the alternative splicing events were determined. In addition to the identification of novel regulated genes, the authors identified multiple transcripts undergoing post-transcriptional regulation. The authors also identified that the circadian rhythm regulates the spliceosome machinery in Arabidopsis by affecting the genes including SPLICING FACTOR 30 (AtSPF30). (Summary by Suresh Damodaran) bioRxiv 10.1101/845560.