Evolution of transposon-encoded anti-silencing factors in Arabidopsis ($)
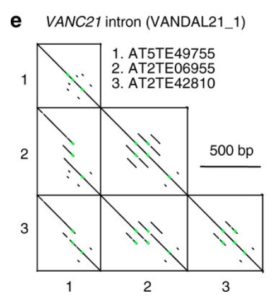 Transposable elements (TEs) are a major component of eukaryotic genomes. Their activity is silenced by epigenetic mechanisms such as chromatin modifications and DNA methylation in order to avoid deleterious effects on host genome stability. Nevertheless, how TEs overcome silencing by the host and propagate is poorly understood. In this study, the authors describe the activity of sequence-specific anti-silencing proteins encoded by Arabidopsis TE families. They show that the anti-silencing VANC21 protein, encoded by members of the TE family VANDAL21, is sufficient for triggering sequence-specific demethylation of VANDAL21 copies. VANC21 binds to specific nine-bases motifs derived from tandem repeats, localized in the non-coding region of VANDAL21. Interestingly, they also identified proteins related to VANC encoded by other TE families, such as the protein VANC6 encoded by VANDAL6 family. VANC6 shows the same behavior as VANC21 and analysis of their target sequences suggests similar evolutionary dynamics. Altogether, this study describes the evolution of TE-specific anti-silencing mechanisms through the behavior of VANC proteins. Such a system could potentially be harnessed to engineer epigenetic states at specific loci, notably in the perspective of genetically modified crops (Summary by Matthias Benoit) Nature Comm. 10.1038/s41467-017-02150-7.
Transposable elements (TEs) are a major component of eukaryotic genomes. Their activity is silenced by epigenetic mechanisms such as chromatin modifications and DNA methylation in order to avoid deleterious effects on host genome stability. Nevertheless, how TEs overcome silencing by the host and propagate is poorly understood. In this study, the authors describe the activity of sequence-specific anti-silencing proteins encoded by Arabidopsis TE families. They show that the anti-silencing VANC21 protein, encoded by members of the TE family VANDAL21, is sufficient for triggering sequence-specific demethylation of VANDAL21 copies. VANC21 binds to specific nine-bases motifs derived from tandem repeats, localized in the non-coding region of VANDAL21. Interestingly, they also identified proteins related to VANC encoded by other TE families, such as the protein VANC6 encoded by VANDAL6 family. VANC6 shows the same behavior as VANC21 and analysis of their target sequences suggests similar evolutionary dynamics. Altogether, this study describes the evolution of TE-specific anti-silencing mechanisms through the behavior of VANC proteins. Such a system could potentially be harnessed to engineer epigenetic states at specific loci, notably in the perspective of genetically modified crops (Summary by Matthias Benoit) Nature Comm. 10.1038/s41467-017-02150-7.