ePlant: Visualizing and exploring multiple levels of data for hypothesis generation ($)
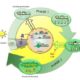
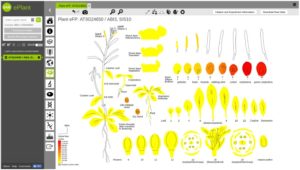 The application of systems biology is quite phenomenal these days for prediction-based modeling and interactive data visualization. Along with the genome sequencing of the model plant Arabidopsis thaliana, there has been a parallel increase in systems biology tools. Unfortunately, these tools have been developed by different groups or institutions for various purposes, whereas a unifying and comprehensive systems biology toolbox is more convenient to connect multiple databases and generate prediction models or build hypothesis. ePlant provides that purpose for Arabidopsis thaliana. In ePlant, Wrase et al. incorporated multiple datasets and integrated search system for analysis and visualization in a single portal. Here, users may access genome, proteome, interactome, transcriptome, sub-cellular localization, ecotype distribution, and 3D molecular structure data. ePlant provides lucid visualization for multiple level of data in a single platform, and can be built upon for research in other species. Plant Cell 10.1105/tpc.17.00073
The application of systems biology is quite phenomenal these days for prediction-based modeling and interactive data visualization. Along with the genome sequencing of the model plant Arabidopsis thaliana, there has been a parallel increase in systems biology tools. Unfortunately, these tools have been developed by different groups or institutions for various purposes, whereas a unifying and comprehensive systems biology toolbox is more convenient to connect multiple databases and generate prediction models or build hypothesis. ePlant provides that purpose for Arabidopsis thaliana. In ePlant, Wrase et al. incorporated multiple datasets and integrated search system for analysis and visualization in a single portal. Here, users may access genome, proteome, interactome, transcriptome, sub-cellular localization, ecotype distribution, and 3D molecular structure data. ePlant provides lucid visualization for multiple level of data in a single platform, and can be built upon for research in other species. Plant Cell 10.1105/tpc.17.00073