CYCLOIDEA3 is targeted by disparate transcription factors in patterning flowers in Gerbera
Yunqing Yu
Donald Danforth Plant Science Center
Kellogg Lab
SAINT LOUIS, MO 63132
United States of America
[email protected]
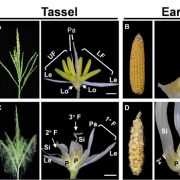
Flowering plants have evolved enormous diversity and complexity of inflorescence architecture. One great model for studying complex inflorescence development and evolution is the Asteraceae (the sunflower family), which is one of the largest flowering plant families and widely distributed across the world. The inflorescence of Asteraceae species is head-like, composed of showy ray flowers in the outer whorls and densely clustered disc flowers at the center. The genus Gerbera has an additional flower type called trans flowers, which are similar to ray flowers but smaller in size (Fig. 1A, B). The ray and trans flowers are zygomorphic (bilaterally symmetrical) and male sterile, with three ventral petals that fuse together and form a showy ligule to attract pollinators, while the other two dorsal petals remain rudimentary. By contrast, disc flowers are actinomorphic (radially symmetrical), with functional carpels and stamens. The petals of a disc flower are fused at the base and separated at the tip, forming a tubular corolla. In all three flower types, the sepals are highly modified into hair-like structures called pappus bristles (Fig. 1A, B) (Tähtiharju et al., 2012).
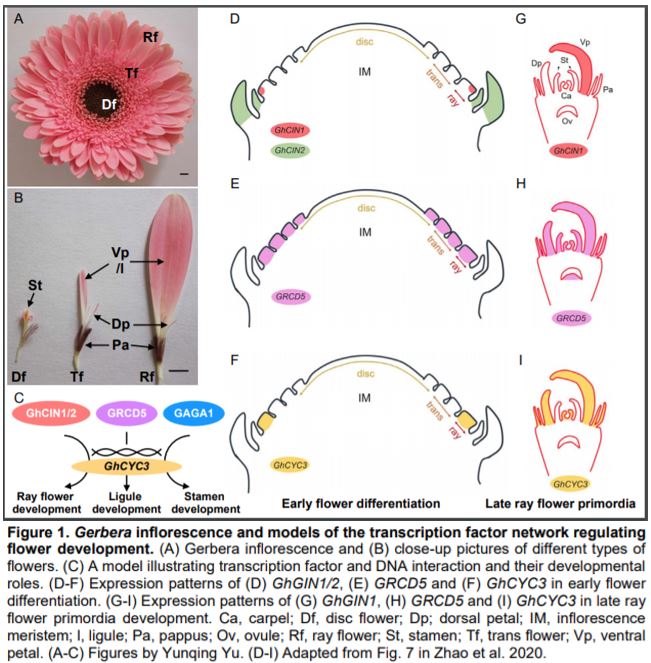
Morphological innovation of inflorescence architecture is often achieved by modifying the ancestral gene regulatory network underlying flower development. In this issue of Plant Physiology, Zhao et al. identify a network of TCP and MADS-box transcription factors (TFs) regulating flower symmetry and identity in Gerbera hybrida. Previous work has shown that the class II TCP TFs CYCLOIDEA (CYC) are required for ray flower symmetry and development in Asteraceae (Broholm et al., 2008; Fambrini et al., 2011). In both sunflower (Helianthus annuus) and Gerbera, CYC2 clade genes are preferentially expressed in ray flower primordia as compared to disc flower primordia (Tähtiharju et al., 2012). The non-functional mutation of sunflower CYC2c transforms ray flowers into partially disc-like flowers, with tubular corollas and fertile reproductive organs (Fambrini et al., 2011). Conversely, overexpression of Gerbera CYC2 causes disc flowers to gain ray flower characteristics (Broholm et al., 2008). Outside of Asteraceae, Antirrhinum majus (snapdragon) CYC and its paralog DICHOTOMA are expressed in the dorsal floral organs and necessary for dorsal/ventral asymmetry, suggesting that CYC clade genes have been recruited independently during the evolution of zygomorphic flowers (Manassero et al., 2013).
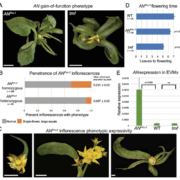
In this issue, Zhao et al. aimed to identify the upstream TFs that directly regulate the transcription of GhCYC3, one of the six CYC2 clade genes in Gerbera. They first identified a 27 bp conserved element in the promoter region of Asteraceae CYC2 clade genes. This element was used as bait in a yeast one-hybrid assay to screen an Arabidopsis thaliana TF library. The authors then analyzed the expression pattern of the Gerbera homologs of 28 candidate Arabidopsis TFs. Ten Gerbera TFs corresponding to five candidate Arabidopsis TFs showed differential expression between flower types and thus were further pursued. Using luciferase reporter assays, two of the candidate TFs, GhCIN1 and GhCIN2 in the class II CIN-like TCP TF clade, activated transcription (Fig. 1C) (Zhao et al., 2020). As an alternative strategy, Zhao et al. also tested if the well-known flower developmental regulators, MADS-box TFs, act upstream of GhCYC3. They showed that the C-class TF GAGA1 and E-class TFs GRCD5/GRCD8 activated the luciferase reporter (Fig. 1C). Zhao et al. also demonstrated that the TCP and MADS-box binding motifs (GGNCC and CArG, respectively) are present in GhCYC3 promoter, and required for the transcriptional activation of the reporter. These results reveal that members of TCP and MADS-box TFs activate GhCYC3 expression.
If these candidate TFs directly regulate GhCYC3 expression in Gerbera, they should share correlated spatiotemporal expression patterns with GhCYC3. At an early developmental stage, GhCYC3 is specifically expressed in ray flower primordia, but not in trans or disc flower primordia (Fig. 1F). At later stages, GhCYC3 transcripts can be detected in petals and carpels in all flower types, but are absent in stamens (Juntheikki‐Palovaara et al., 2014) (Fig. 1I). GhCIN1 and GhCIN2 also have higher expression in ray flower primordia than in disc flower primordia. However, GhCIN1 is restricted to the ventral side of ray flower primordia and later in ventral petals, while GhCIN2 is mainly expressed in bracts (Fig. 1D, G). Previous study has shown broad expression of GRCDs in the primordia of all three flower types and reproductive organs (Fig. 1E, H) (Zhang et al., 2017). Here, Zhao et al. demonstrated similar expression trends of GRCD5 and GhCYC3 in ventral petal (ligule) development. GAGA1 has a classic C-class TF expression pattern in stamens and carpels regardless of flower type (Yu et al., 1999). These results reveal overlapped but not identical expression patterns between GhCYC3 and its upstream TFs, suggesting complex regulation of GhCYC3 expression by multiple TFs.
Last but not least, genetic studies were used to validate the functional relationships of these TFs. Ectopic expression of GhCYC3 affects ligule elongation in disc flowers, and transforms disc flowers into trans- and ray-like flowers (Juntheikki‐Palovaara et al., 2014). RNAi lines of GhCIN1, GhCIN2 and GRCD5 all showed reduced expression of GhCYC3. Compared to the wild-type, the Ghcin1 and Ghcin2 RNAi lines tend to accelerate development of ray flower primordia, although no obvious phenotype was observed in mature flowers (Zhao et al., 2020). Silencing GRCD5 results in defects in petal development, with petals showing greenish and bract-like structures (Zhang et al., 2017).
Combining the above analyses, GhCYC3 is probably regulated by multiple TFs in different floral organs and at different developmental stages. GhCYC3 may be a target of GhCIN1/2 in ray flower primordia development; GRCD5 may activate GhCYC3 in ligule elongation; and GAGA1 may target GhCYC3 in stamen development (Fig. 1C). Considering that all these TFs are in multi-member families and share redundant roles, functional analysis of higher order mutants may further reveal their roles and relationships in flower patterning in Gerbera.
Yunqing Yu
Donald Danforth Plant Science Center
Saint Louis, MO, USA
Reference
Broholm SK, Tähtiharju S, Laitinen RA, Albert VA, Teeri TH, Elomaa P (2008) A TCP domain transcription factor controls flower type specification along the radial axis of the Gerbera (Asteraceae) inflorescence. Proceedings of the National Academy of Sciences 105: 9117-9122
Fambrini M, Salvini M, Pugliesi C (2011) A transposon-mediate inactivation of a CYCLOIDEA-like gene originates polysymmetric and androgynous ray flowers in Helianthus annuus. Genetica 139: 1521-1529
Juntheikki‐Palovaara I, Tähtiharju S, Lan T, Broholm SK, Rijpkema AS, Ruonala R, Kale L, Albert VA, Teeri TH, Elomaa P (2014) Functional diversification of duplicated CYC2 clade genes in regulation of inflorescence development in Gerbera hybrida (Asteraceae). The Plant Journal 79: 783-796
Manassero NGU, Viola IL, Welchen E, Gonzalez DH (2013) TCP transcription factors: architectures of plant form. Biomolecular concepts 4: 111-127
Tähtiharju S, Rijpkema AS, Vetterli A, Albert VA, Teeri TH, Elomaa P (2012) Evolution and diversification of the CYC/TB1 gene family in Asteraceae—a comparative study in Gerbera (Mutisieae) and sunflower (Heliantheae). Molecular Biology and Evolution 29: 1155-1166
Yu D, Kotilainen M, Pöllänen E, Mehto M, Elomaa P, Helariutta Y, Albert VA, Teeri TH (1999) Organ identity genes and modified patterns of flower development in Gerbera hybrida (Asteraceae). The Plant Journal 17: 51-62
Zhang T, Zhao Y, Juntheikki I, Mouhu K, Broholm SK, Rijpkema AS, Kins L, Lan T, Albert VA, Teeri TH (2017) Dissecting functions of SEPALLATA‐like MADS box genes in patterning of the pseudanthial inflorescence of Gerbera hybrida. New Phytologist 216: 939-954
Zhao Y, Broholm SK, Wang F, Rijpkema AS, Lan T, Albert V, Teeri TH, Elomaa P (2020) TCP and MADS-box transcription factor networks regulate heteromorphic flower type identity in Gerbera hybrida. Plant Physiology https://doi.org/10.1104/pp.20.00702