A Lipid Synthesis Enzyme Confers Freezing Tolerance
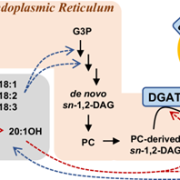
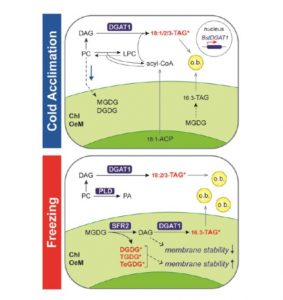 Despite major advances in understanding cold signaling, cold acclimation, and freezing protection in model and crop species, and extensive studies of natural variation in freezing tolerance in Arabidopsis accessions, the question of which genes and mechanisms underlie freezing tolerance of wild species has remained largely unanswered. Because freezing tolerance generally exhibits continuous variation, it is amenable to quantitative trait locus (QTL) mapping to identify relevant chromosomal regions. Arisz et al. (10.1104/pp.18.00503) have performed a quantitative trait locus (QTL) study of acclimated freezing tolerance in seedlings of Boechera stricta, a relative of Arabidopsis that is native to the Rocky Mountains. A single QTL was identified that contained the gene encoding ACYL-COA:-DIACYLGLYCEROL ACYLTRANSFERASE1 (BstDGAT1), whose expression is highly cold responsive. DGAT1 catalyzes the final step in assembly of triacylglycerol (TAG) by acyl transfer from acyl-CoA to diacylglycerol. Freezing tolerant plants showed higher DGAT1 expression during cold acclimation than more sensitive plants, and this resulted in increased accumulation of TAG in response to subsequent freezing. The levels of oligogalactolipids that are produced by SFR2 (SENSITIVE TO FREEZING2), an indispensable element of freezing tolerance in Arabidopsis, were also higher in freezing-tolerant plants. SFR2 transfers galactosyl groups from monogalactosyldiacylglycerol to other galactolipids to generate oligogalactolipids, including tri- and tetragalactosyldiacylglycerol, which are thought to stabilize chloroplast membranes, and diacylglycerol. Overexpression of AtDGAT1 in Arabidopsis also resulted in a substantial increase in freezing tolerance. Together, the data suggest a mechanism by which DGAT1-mediated TAG assembly confers tolerance to freezing stress.
Despite major advances in understanding cold signaling, cold acclimation, and freezing protection in model and crop species, and extensive studies of natural variation in freezing tolerance in Arabidopsis accessions, the question of which genes and mechanisms underlie freezing tolerance of wild species has remained largely unanswered. Because freezing tolerance generally exhibits continuous variation, it is amenable to quantitative trait locus (QTL) mapping to identify relevant chromosomal regions. Arisz et al. (10.1104/pp.18.00503) have performed a quantitative trait locus (QTL) study of acclimated freezing tolerance in seedlings of Boechera stricta, a relative of Arabidopsis that is native to the Rocky Mountains. A single QTL was identified that contained the gene encoding ACYL-COA:-DIACYLGLYCEROL ACYLTRANSFERASE1 (BstDGAT1), whose expression is highly cold responsive. DGAT1 catalyzes the final step in assembly of triacylglycerol (TAG) by acyl transfer from acyl-CoA to diacylglycerol. Freezing tolerant plants showed higher DGAT1 expression during cold acclimation than more sensitive plants, and this resulted in increased accumulation of TAG in response to subsequent freezing. The levels of oligogalactolipids that are produced by SFR2 (SENSITIVE TO FREEZING2), an indispensable element of freezing tolerance in Arabidopsis, were also higher in freezing-tolerant plants. SFR2 transfers galactosyl groups from monogalactosyldiacylglycerol to other galactolipids to generate oligogalactolipids, including tri- and tetragalactosyldiacylglycerol, which are thought to stabilize chloroplast membranes, and diacylglycerol. Overexpression of AtDGAT1 in Arabidopsis also resulted in a substantial increase in freezing tolerance. Together, the data suggest a mechanism by which DGAT1-mediated TAG assembly confers tolerance to freezing stress.