Small Secreted Peptides of a Model Legume: a Database
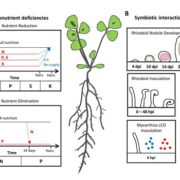
Plant small secreted peptides (SSPs) have emerged as an important class of regulatory molecules involved in plant growth, development, 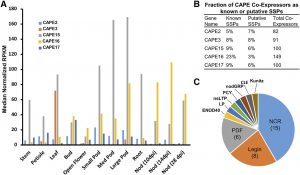 plant-microbe interactions, and stress tolerance. SSPs are typically encoded within preproteins of 100–250 amino acids, that are subsequently processed into shorter bioactive peptides of ~5 to 50 residues that act at very low, often nanomolar physiological concentrations. SSP-coding genes are frequently overlooked, because genome annotation pipelines generally ignore small open reading frames, which are those most likely to encode SSPs. Also, SSP-coding small open reading frames are often expressed at low levels or only under specific conditions, and thus are underrepresented in non-tissue-targeted or non-condition-optimized RNA-sequencing projects. SSPs are of particular significance for legumes, since recent discoveries show that SSPs regulate symbiotic root nodulation and root development. Boschiero et al. (10.1104/pp.19.01088) previously identified 4,439 SSP-encoding genes in the model legume Medicago truncatula. To support systematic characterization and annotation of these putative SSP-encoding genes, they have developed the M. truncatula Small Secreted Peptide Database (MtSSPdb). MtSSPdb currently hosts (1) a compendium of M. truncatula SSP candidates with putative function and family annotations; (2) a large-scale M. truncatula RNA-sequencing-based gene expression atlas integrated with various analytical tools, including differential expression, co-expression, and pathway enrichment analyses; (3) an online plant SSP prediction tool capable of analyzing protein sequences at the genome scale using the same protocol as for the identification of SSP genes; and (4) information about a library of synthetic peptides and root and nodule phenotyping data from synthetic peptide screens in planta. MtSSPdb is an important resource for the plant scientific community and has the potential to become the most complete database of SSPs in plants.
plant-microbe interactions, and stress tolerance. SSPs are typically encoded within preproteins of 100–250 amino acids, that are subsequently processed into shorter bioactive peptides of ~5 to 50 residues that act at very low, often nanomolar physiological concentrations. SSP-coding genes are frequently overlooked, because genome annotation pipelines generally ignore small open reading frames, which are those most likely to encode SSPs. Also, SSP-coding small open reading frames are often expressed at low levels or only under specific conditions, and thus are underrepresented in non-tissue-targeted or non-condition-optimized RNA-sequencing projects. SSPs are of particular significance for legumes, since recent discoveries show that SSPs regulate symbiotic root nodulation and root development. Boschiero et al. (10.1104/pp.19.01088) previously identified 4,439 SSP-encoding genes in the model legume Medicago truncatula. To support systematic characterization and annotation of these putative SSP-encoding genes, they have developed the M. truncatula Small Secreted Peptide Database (MtSSPdb). MtSSPdb currently hosts (1) a compendium of M. truncatula SSP candidates with putative function and family annotations; (2) a large-scale M. truncatula RNA-sequencing-based gene expression atlas integrated with various analytical tools, including differential expression, co-expression, and pathway enrichment analyses; (3) an online plant SSP prediction tool capable of analyzing protein sequences at the genome scale using the same protocol as for the identification of SSP genes; and (4) information about a library of synthetic peptides and root and nodule phenotyping data from synthetic peptide screens in planta. MtSSPdb is an important resource for the plant scientific community and has the potential to become the most complete database of SSPs in plants.