Multiple Phytohormone Screening Method
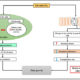
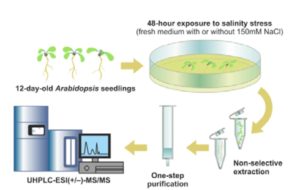 Phytohormones are naturally occurring signaling molecules that play key roles in the regulation of plant physiology, development, and adaptation to environmental stimuli. Generally, their concentrations in plant tissues are extremely low (fmol-pmol/g fresh weight, FW). Although certain phytohormones are usually related to specific biological functions or responses, there is increasing evidence that plant hormone signaling involves complex interactions (‘crosstalk’) among all the pathways involved. Clearly, a convenient method to simultaneously quantify a wide range of plant signaling molecules greatly facilitate the investigation of hormone functions and networks. In this issue, Šimura et al. (10.1104/pp.18.00293) present a method for the simultaneous targeted profiling of phytohormone-related analytes from minute amounts of fresh plant material (< 20 mg). Rapid and non-selective extraction, fast one-step sample purification, and extremely sensitive ultra-high-performance liquid chromatography-tandem mass spectrometry (UHPLC-MS/MS) enable concurrent quantification of the main phytohormone classes: cytokinins, auxins, brassinosteroids, gibberellins, jasmonates, salicylates, and abscisates. To assess the practical utility of this ‘hormonomic’ approach, the method was applied to characterize phytohormone profiles in root and shoot tissues of control and salt-stressed Arabidopsis (Arabidopsis thaliana) seedlings. The authors successfully quantified a total of 43 endogenous compounds in both root and shoot samples. These findings suggest that physiological responses to stimuli such as salinity are not controlled solely by a single active form of a hormone (or even single active forms of several hormones), but by the combined activities and ratios of multiple hormones, metabolites, and (hence) genes.
Phytohormones are naturally occurring signaling molecules that play key roles in the regulation of plant physiology, development, and adaptation to environmental stimuli. Generally, their concentrations in plant tissues are extremely low (fmol-pmol/g fresh weight, FW). Although certain phytohormones are usually related to specific biological functions or responses, there is increasing evidence that plant hormone signaling involves complex interactions (‘crosstalk’) among all the pathways involved. Clearly, a convenient method to simultaneously quantify a wide range of plant signaling molecules greatly facilitate the investigation of hormone functions and networks. In this issue, Šimura et al. (10.1104/pp.18.00293) present a method for the simultaneous targeted profiling of phytohormone-related analytes from minute amounts of fresh plant material (< 20 mg). Rapid and non-selective extraction, fast one-step sample purification, and extremely sensitive ultra-high-performance liquid chromatography-tandem mass spectrometry (UHPLC-MS/MS) enable concurrent quantification of the main phytohormone classes: cytokinins, auxins, brassinosteroids, gibberellins, jasmonates, salicylates, and abscisates. To assess the practical utility of this ‘hormonomic’ approach, the method was applied to characterize phytohormone profiles in root and shoot tissues of control and salt-stressed Arabidopsis (Arabidopsis thaliana) seedlings. The authors successfully quantified a total of 43 endogenous compounds in both root and shoot samples. These findings suggest that physiological responses to stimuli such as salinity are not controlled solely by a single active form of a hormone (or even single active forms of several hormones), but by the combined activities and ratios of multiple hormones, metabolites, and (hence) genes.