It’s a TRAPP! Arabidopsis Transport Protein Particle (TRAPP) Complexes Contain a Novel Plant-Specific Subunit
Membrane vesicle trafficking is the process of moving and distributing signaling molecules from their place of synthesis or capture to target locations inside or outside the cell. Vesicles, mobile membrane compartments produced from the endoplasmic reticulum, carry proteins and other molecules to an acceptor compartment. This is a complex process, with many accessory proteins required for proper vesicle formation, migration, tethering, and acceptor fusion. Transport Protein Particle (TRAPP) complexes are multiunit proteins that have a critical role in trafficking, where they act as guanine nucleotide exchange factors (GEF) for Ypt/Rab GTPases mediating protein transport. Composed of multiple, highly-conserved subunits, two TRAPP complexes (TRAPPII and TRAPPIII) have been identified in yeast and in animals (Barrowman et al., 2010; Reidel et al. 2018). However, to date relatively little is known about TRAPP complexes in plant systems (Kalde, et al. 2019; Rosquete et al., 2019).
 Here, Garcia et al. (2020) investigated key features of Arabidopsis TRAPP complexes and characterized plant-specific TRAPP subunits. First, the authors used transgenic knockout mutants to examine the role played by TRS33—a protein essential for the membrane association of TRAPPII-specific subunits—on the TRAPPII complex. They examined whether trs33-1 mutants showed altered localization of the TRAPPII-specific subunit AtTRS120 and found that TRS120 localizes to the cell plate only in the presence of AtTRS33, suggesting that TRS33 is essential for proper localization of TRAPPII complexes during cytokinesis. The authors then used a multistage protocol combining stable isotope labelling in planta with mass spectroscopy (SILIP-MS; Schaff et al., 2008) to identify proteins associated with AtTRS33. Fourteen proteins were identified as interactors of TRS33, including orthologs of six proteins found in all TRAPP complexes, seven orthologs of complex-specific subunits, and one novel protein, which the authors named TRAPP-interacting plant protein (TRIPP). Next, the authors repeated the SILIP-MS protocol using TRIPP as bait and found that only the shared and TRAPPII-specific subunits were pulled down by TRIPP, suggesting that the plant-specific TRIPP subunit is present only in TRAPPII complexes. Interestingly, the authors found that TRIPP interacts with two TRAPPII-specific subunits and localizes to the Trans-Golgi Network/early endosome (TGN/EE), a dynamic hub of vesicle-mediated trafficking.
Here, Garcia et al. (2020) investigated key features of Arabidopsis TRAPP complexes and characterized plant-specific TRAPP subunits. First, the authors used transgenic knockout mutants to examine the role played by TRS33—a protein essential for the membrane association of TRAPPII-specific subunits—on the TRAPPII complex. They examined whether trs33-1 mutants showed altered localization of the TRAPPII-specific subunit AtTRS120 and found that TRS120 localizes to the cell plate only in the presence of AtTRS33, suggesting that TRS33 is essential for proper localization of TRAPPII complexes during cytokinesis. The authors then used a multistage protocol combining stable isotope labelling in planta with mass spectroscopy (SILIP-MS; Schaff et al., 2008) to identify proteins associated with AtTRS33. Fourteen proteins were identified as interactors of TRS33, including orthologs of six proteins found in all TRAPP complexes, seven orthologs of complex-specific subunits, and one novel protein, which the authors named TRAPP-interacting plant protein (TRIPP). Next, the authors repeated the SILIP-MS protocol using TRIPP as bait and found that only the shared and TRAPPII-specific subunits were pulled down by TRIPP, suggesting that the plant-specific TRIPP subunit is present only in TRAPPII complexes. Interestingly, the authors found that TRIPP interacts with two TRAPPII-specific subunits and localizes to the Trans-Golgi Network/early endosome (TGN/EE), a dynamic hub of vesicle-mediated trafficking.
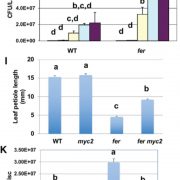
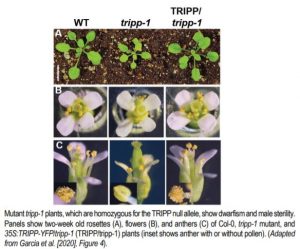
Garcia and colleagues also found important phenotypic abnormalities in plants homozygous for a null TRIPP allele (caused by a T-DNA insertion in the fifth exon), but not for plants heterozygous for this allele (see Figure). Despite producing superficially normal anthers and stigma, homozygous tripp-1 mutants were sterile, with anthers producing no pollen and ovules producing no seed when fertilized by wild-type pollen. TRIPP mutations were also associated with dwarfism, as five-week-old plants homozygous for the null allele were significantly shorter than wild type plants. Moreover, tripp-1 plants also showed differences in skotomorphogenic growth. When growth in the dark, homozygous mutant plants showed shorter hypocotyl length and longer root length than Col-0 wild type plants, but these phenotypes were not observed when plants were grown in the light. This phenotype may be linked to auxin signaling, since tripp-1 mutants also show defective polar localization of PIN-FORMED 2 (PIN2), an important auxin transporter.
Taken together, these results improve our understanding of plant TRAPP complexes. The authors show that TRAPPII (and possibly TRAPPIII) complexes are present in plants, and that plant TRAPPII complexes contain TRIPP, a novel, plant-specific TRAPP component. Moreover, the authors were also able to show that TRIPP is implicated in plant growth and development, presumably by way of altering TRAPPII membrane association. Future studies of the role TRIPP plays in plant TRAPPII complex function may shed light on how TRAPP complexes in plants differ from those in other metazoans.
William Hughes
Department of Ecology, Environment, and Plant Sciences,
Stockholm University
Svante Arrhenius väg 20
114 18 Stockholm
ORCID: 0000-0003-4142-5279
REFERENCES
Barrowman, J., Bhandari, D., Reinisch, K., and Ferro-Novick, S. (2010). TRAPP complexes in membrane traffic: convergence through a common Rab. Nature Rev. Molec. Cell Biol. 11: 759-763. DOI: 10.1038/nrm2999
Garcia, V., Xu, S., Ravikumar, R., Wang, W., Elliott, L., Gonzalez, E., Fesenko, M., Altmann, M., Brunschwieger, B., Falter-Braun, P., Moore, I., Burlingame, A., Assaad, F., and Wang, Z. (2020). Proteomic studies of the Arabidopsis TRAPP complexes reveal conserved organization and a novel plant-specific component with a role in plant development. Plant Cell. Published April 2020. DOI: https://doi.org/10.1105/tpc.20.00044.
Kalde, M., Elliott, L., et al. (2019). Interactions between Transport Protein Particle (TRAPP) complexes and Rab GTPases in Arabidopsis. Plant J. 100: 279-297. DOI: 10.1111/tpj.14442.
Riedel, F., Galindo, A., Muschalik, N., and Munro, S. (2018). The two TRAPP complexes of metazoans have distinct roles and act on different Rab GTPases. J. Cell Biol. 217: 601-617. DOI: 10.1083/jcb.201705068.
Rosquete, M., Worden, N., Ren, G., Sinclair, R., Pfleger, S., Salemi, M., Phinney, B., Domozych, D., Wilkop, T., and Drakakaki, G. (2019) AtTRAPP11/ROG2: A role for TRAPPs in maintenance of the Plant Trans-Golgi network/early endosome organization and function. Plant Cell. 31: 1879-1898. DOI: 10.1105/tpc.19.00110
Schaff, J. E., Mbeunkui, F., Blackburn, K., Bird, D. M., and Goshe, M. B. (2008). SILIP: a novel stable isotope labeling method for in planta quantitative proteomic analysis. Plant J. 56: 840-854. DOI: 10.1111/j.1365-313X.2008.03639.x