Invisible No Longer: Peptidoglycan in Moss Chloroplasts
IN BRIEF by Nancy Hofmann [email protected]
Most bacteria have a peptidoglycan layer between the inner and outer membranes (reviewed in Typas et al., 2012). The cyanobacterial endosymbiont that gave rise to plastids would have contained such a peptidoglycan wall including d-amino acids. Indeed, peptidoglycan can be visualized by electron microscopy in the envelope of cyanelles, also known as muroplasts—the plastids of glaucophytes. However, peptidoglycan has not been observed in the envelopes of plastids of other lineages, suggesting that it has been lost in the course of plastid evolution (reviewed in Takano and Takechi, 2010). Surprisingly, though, antibiotics that target peptidoglycan biosynthesis cause plastid phenotypes in species that do not have a visible peptidoglycan layer in electron micrographs. For instance, d-cycloserine induces macrochloroplasts in the moss Physcomitrella patens. Furthermore, some algae and plants, including P. patens, have the full complement of genes for peptidoglycan biosynthesis. Disruption of such Mur genes leads to macrochloroplast formation in P. patens. This apparent connection between peptidoglycan and chloroplast division prompted Hirano et al. (2016) to revisit the question of whether land plant plastids contain a peptidoglycan layer.
d-Cycloserine inhibits d-Ala:d-Ala ligases (DDLs), which produce d-Ala-d-Ala (DA-DA) during peptidoglycan biosynthesis in bacteria. Based on this, Hirano et al. characterized DDL in P. patens, showing that it is targeted to the chloroplast and that knockout ddl mutants have macrochloroplasts similar to those induced by d-cycloserine. The moss ddl mutants had increased levels of d-Ala, consistent with accumulation of the substrate in the absence of DDL activity. Importantly, exogenous DA-DA could partially rescue the macrochloroplast phenotype of the mutants. These data again pointed to the likely importance of peptidoglycan in plastid division in moss.
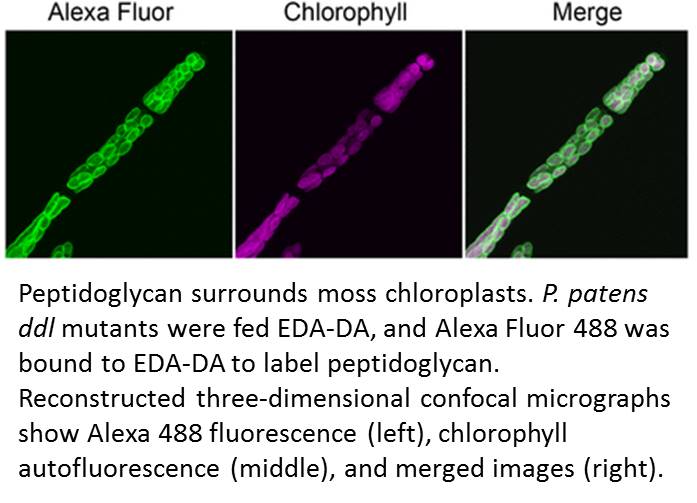
Accordingly, Hirano et al. made a renewed attempt to visualize peptidoglycan in a land plant. They used a metabolic labeling technique that recently revealed peptidoglycan in Chlamydia, which had been so recalcitrant to peptidoglycan visualization that the phenomenon was termed the Chlamydial anomaly (Liechti et al., 2014). Hirano and coworkers fed the P. patens ddl mutant EDA-DA, an analog of DA-DA that was able to rescue the chloroplast phenotype as well as DA-DA. After the cells were fixed, the authors used click chemistry to link an azide-modified Alexa Fluor to the alkyne functional group of EDA-DA. Excitingly, this approach revealed a layer surrounding each chloroplast (see figure) and forming at the division plane of dividing chloroplasts.
This Breakthrough Report from Hirano et al. establishes that peptidoglycan is present in the plastids of at least some land plants and is needed for chloroplast division in P. patens. The role of peptidoglycan in chloroplast division does not appear to be conserved in Arabidopsis thaliana, which does not have all of the genes for peptidoglycan biosynthesis. Thus, it remains likely that peptidoglycan has been lost, likely multiple times, over the course of evolution of plastid-containing lineages. However, this work emphasizes that the lack of a visible peptidoglycan layer in electron micrographs cannot be taken to reflect the absence of peptidoglycan in vivo.
Hirano, T., Tanidokoro, K., Shimizu, Y., Kawarabayasi, Y., Ohshima, T., Sato, M., Tadano, S., Ishikawa, H., Takio, S., Takechi, K., Takano, H. (2016). Moss chloroplasts are surrounded by a peptidoglycan wall containing d-amino acids. Plant Cell 28: 1521–1532.
Liechti, G.W., Kuru, E., Hall, E., Kalinda, A., Brun, Y.V., VanNieuwenhze, M., Maurelli, A.T. (2014). A new metabolic cell-wall labelling method reveals peptidoglycan in Chlamydia trachomatis. Nature 506: 507–510.
Takano, H., Takechi, K. (2010). Plastid peptidoglycan. Biochim. Biophys. Acta 1800: 144–151.
Typas, A., Banzhaf, M., Gross, C.A., Vollmer, W. (2012). From the regulation of peptidoglycan synthesis to bacterial growth and morphology. Nat. Rev. Microbiol. 10: 123–136.







Leave a Reply
Want to join the discussion?Feel free to contribute!