Insights into Salicylic Acid and Mitochondria
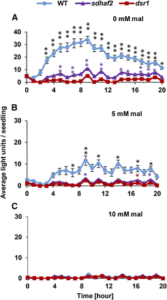 Within the mitochondrial electron transport chain, complex II (succinate dehydrogenase [SDH]) oxidizes succinate to fumarate by transferring electrons to ubiquinone (UQ), which is reduced to ubiquinol. The enzyme is formed by four subunits: a flavoprotein (SDH1), which contains the FAD cofactor, an iron sulfur (Fe-S) protein (SDH2) housing three Fe-S clusters, and two small integral membrane proteins (SDH3 and SDH4), anchoring the enzyme to the inner membrane and forming the UQ binding site. Several assembly factors have been identified that facilitate FAD and Fe-S insertion into SDH subunits, and one of these, SDHAF2, has been characterized in Arabidopsis. Complexes I and III have been long considered to be the major sources of reactive oxygen species (ROS) production inside mitochondria (mtROS), but recent studies in both mammals and plants have demonstrated that complex II can also be a significant source of mtROS. The knockout of the SDH complex or its assembly factors in plants is lethal, largely preventing its direct study through gene deletion in plants. This limitation changed when a point mutation of SDH1-1 (disrupted in stress response1 [dsr1]) was identified that did not knockout SDH, but instead lowered SDH activity and decreased mitochondrial ROS production. The dsr1 mutant exhibits a reduced expression of a set of salicylic acid (SA)-responsive genes normally induced in response to exposure of Arabidopsis to bacterial, fungal, or viral pathogens. The dsr1 mutant also had higher susceptibility to fungal and bacterial pathogens, indicating that mitochondrial SDH is involved in response to biotic stress in vivo in plants. Belt et al. () have further analyzed the role of mitochondrial SDH in SA signaling using dsr1 and sdhaf2 mutants. Both mutants showed strongly decreased SA-inducible stress promoter responses and low SDH maximum capacity compared to the wild type. Both dsr1 and sdhaf2 showed lower rates of SA-dependent H2O2 production in vitro in keeping with their low SA-dependent stress signaling responses in vivo. The authors provide quantitative and kinetic evidence that SA acts at or near the ubiquinone binding site of SDH to stimulate activity and contributes to plant stress signaling by increased rates of mitochondrial H2O2 production, leading to part of the SA-dependent transcriptional response in plant cells.
Within the mitochondrial electron transport chain, complex II (succinate dehydrogenase [SDH]) oxidizes succinate to fumarate by transferring electrons to ubiquinone (UQ), which is reduced to ubiquinol. The enzyme is formed by four subunits: a flavoprotein (SDH1), which contains the FAD cofactor, an iron sulfur (Fe-S) protein (SDH2) housing three Fe-S clusters, and two small integral membrane proteins (SDH3 and SDH4), anchoring the enzyme to the inner membrane and forming the UQ binding site. Several assembly factors have been identified that facilitate FAD and Fe-S insertion into SDH subunits, and one of these, SDHAF2, has been characterized in Arabidopsis. Complexes I and III have been long considered to be the major sources of reactive oxygen species (ROS) production inside mitochondria (mtROS), but recent studies in both mammals and plants have demonstrated that complex II can also be a significant source of mtROS. The knockout of the SDH complex or its assembly factors in plants is lethal, largely preventing its direct study through gene deletion in plants. This limitation changed when a point mutation of SDH1-1 (disrupted in stress response1 [dsr1]) was identified that did not knockout SDH, but instead lowered SDH activity and decreased mitochondrial ROS production. The dsr1 mutant exhibits a reduced expression of a set of salicylic acid (SA)-responsive genes normally induced in response to exposure of Arabidopsis to bacterial, fungal, or viral pathogens. The dsr1 mutant also had higher susceptibility to fungal and bacterial pathogens, indicating that mitochondrial SDH is involved in response to biotic stress in vivo in plants. Belt et al. () have further analyzed the role of mitochondrial SDH in SA signaling using dsr1 and sdhaf2 mutants. Both mutants showed strongly decreased SA-inducible stress promoter responses and low SDH maximum capacity compared to the wild type. Both dsr1 and sdhaf2 showed lower rates of SA-dependent H2O2 production in vitro in keeping with their low SA-dependent stress signaling responses in vivo. The authors provide quantitative and kinetic evidence that SA acts at or near the ubiquinone binding site of SDH to stimulate activity and contributes to plant stress signaling by increased rates of mitochondrial H2O2 production, leading to part of the SA-dependent transcriptional response in plant cells.



Leave a Reply
Want to join the discussion?Feel free to contribute!