Identifying an Unexpected Player in DNA Methylation
Choudry et al. show that the mRNA export protein ALY1 enables genome-wide RNA-directed DNA methylation, transposon repression and transgene silencing though the efficient export of the AGO6 mRNA. Plant Cell https://doi.org/10.1105/tpc.18.00624
By Sarah Choudury
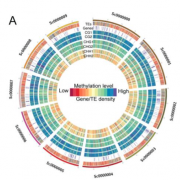
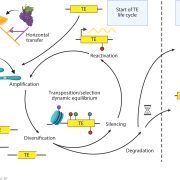
Background: Transposons, sometimes called “jumping genes,” are mobile genetic elements that make up large portions of plant and animal genomes. When a transposon jumps to a new place in the genome, it can cause mutations by disrupting the region it enters. Because of this potential to cause harm, mechanisms have evolved to repress transposon activity. In plants, these mechanisms include addition of repressive marks such as DNA and histone methylation at transposon loci, which act to “turn off” their activity. These repressive marks are targeted to transposons through the action of small interfering RNAs (siRNAs) and proteins in a process called “RNA-directed DNA methylation” (RdDM). While many of the proteins and RNAs required for RdDM have been identified, it is a complex process with many layers of regulation that we are still trying to understand.
Question: We set out to identify new proteins required for RdDM and transposon repression. To do this, we screened Arabidopsis mutants looking for ones that had reduced DNA methylation at transposon loci, indicating they were defective for RdDM.
Findings: We identified an Arabidopsis mutant that had reduced transposon DNA methylation at thousands of transposons throughout the genome. The mutation that caused this loss of DNA methylation was in the ALY1 gene. It has been previously shown that ALY1 exports many messenger RNAs (mRNAs) from the nucleus to the cytoplasm, where they can then be translated into proteins. Our work linked the role of ALY1 in mRNA transport and its newly identified role in DNA methylation by showing that ALY1 exports an mRNA that codes for the critical RdDM protein ARGONAUTE6 (AGO6). In the absence of ALY1, AGO6 mRNA is not properly exported from the nucleus to the cytoplasm, leading to a reduction in AGO6 protein accumulation and ultimately reduced DNA methylation at transposons.
Next steps: Future studies include screens to identify other factors required for RdDM. It will also be of interest to pinpoint which factors are necessary and/or sufficient for different phases of RdDM such as initiation, establishment, and the transition to other silencing pathways.
Sarah G. Choudury, Saima Shahid, Diego Cuerda-Gil, Kaushik Panda, Alissa Cullen, Qurat Ul Ayn Ashraf, Meredith J. Sigman, Andrea D. McCue and R. Keith Slotkin. (2019). The RNA Export Factor ALY1 Enables Genome-Wide RNA-Directed DNA Methylation. Plant Cell. https://doi.org/10.1105/tpc.18.00624
Key words: DNA methylation, mRNA export, transposons