How can genomics help neglected crops fight disease?
Guest post by Kelsey Wood (@klsywd) a PhD student researching the genetics and genomics of plant-pathogen interactions at the University of California, Davis.
I recently attended a Plant Pathology symposium on “Genomics Strategies for Developing Sustainable Disease Resistance for Neglected Crops in the Developing World“. The symposium was held at the University of California, Berkeley and was hosted by the Innovative Genomics Institute (IGI) and the Open Philanthropy Project.
As it is only a ~1 hour drive from Davis, myself and 10 others from the Michelmore lab woke up early to commute to Berkeley for the symposium. There was a great lineup of speakers that kept the audience engaged for the day-long event.
As is my habit, instead of taking notes by pen and paper or on my Evernote, I used Twitter to simultaneously take notes and share the information with my followers and the greater Twitter-verse. I used the hashtag #PlantPathIGI to tag all of my tweets related to the conference so myself and others could easily find them.
Since many of my followers are fellow plant geneticists and plant pathologists, my tweets received a lot of attention from scientists and others across the world. This is one of my favorite things about Twitter, the ability to have real time conversations and sharing of information across geographical barriers!
For those of you who may have missed it on Twitter, I will give a summary of the conference highlights in this blog post.
The first talk was by Diana Horvath from the 2Blades Foundation, which aims to bridge the gap between basic research into the molecular basis of plant-pathogen interactions and the translational applications. They work directly with research scientists at The Sainsbury Lab and other institutions to bring discoveries to the field. She discussed several of their projects on wheat rusts, Asian soybean rust, and cassava brown streak. Their main strategy, which turned out to be a theme for most of the symposium, is the stacking of several resistance genes (‘R genes’) to confer more durable resistance than a single R gene alone.
The mission of @2blades – effective and durable plant disease resistance, using R genes -Diana Horvath #PlantPathIGI @igisci
— kelsey wood (@klsywd) January 25, 2017
Next up was Jagger Harvey of Kansas State on “Bringing Orphan Crops into the Genetic Family“. He started off by suggesting that increasing agricultural productivity to feed the 8.3 billion people projected to be in the world by 2030 was not enough – that we also need to prevent post-harvest loss of our food due to disease and contamination by mycotoxins such as aflatoxin. He also highlighted the importance of so-called “Orphan Crops”, which are non-staple crops that may not provide the majority of calories but are valuable culturally and for their micronutrients. Micronutrients include Vitamin A, iron, and zinc and are critical to prevent stunting and developmental delays which are a major problem facing children in Africa and elsewhere in the developing world.
8.3 billion ppl by 2030 – how can we make enough food? Boost production (obv) but also stop post harvest loss –@PHLILDirector #PlantPathIGI
— kelsey wood (@klsywd) January 25, 2017
The next two speakers, Jeff Ellis (CSIRO) and Barbara Valent (Kansas State), both talked about diseases affecting the staple crop wheat. Jeff Ellis talked about wheat rusts, which are well studied and for which several resistance mechanisms are known, including both specific resistance (via R genes) and more broad spectrum resistance (via quantitative resistance genes). Despite all we know, due to rapid evolution of the fungi we need to discover new R genes for breeders to continue to keep up battling rust diseases. While rice is not affected by rust (the reason why is still unknown and this was brought up several times during the symposium), it is greatly affected by rice blast caused by the fungus Magnaporthe oryzae. Interestingly, different pathotypes of this same species can infect wheat and can cause serious damage in specific parts of the world. Barbara Valent demonstrated the importance of the disease in Bangladesh and elsewhere and noted that it is a neglected disease of wheat breeders compared to other threats such as rust. It was clear from both of these talks that plant pathologists and breeders will need to consider multiple diseases when breeding crops for durable resistance, especially in the face of an unpredictable climate.
We need more novel R genes – both NBLRR and non, for wheat rust resistance -Jeff Ellis #PlantPathIGI
— kelsey wood (@klsywd) January 25, 2017
Jonathan Jones of The Sainsbury Lab gave a wonderful talk on how we can use genetics to help reduce crop losses to disease. He noted that genetics is not a “magic bullet” – rather it is one tool of many (including reducing food waste, better agronomy and cultural practices) that we can use to help reach global food security. He talked about the success story of Rpi-vnt1, which is a potato late blight resistance gene and the first to be commercialized just last year. He also emphasized the importance of gene stacking and also the study of basic molecular biology (in his case – the basis of non-host resistance) to discover new resistance genes.
Genetics not a magic bullet but is an important component of the “wedge” for world food security –@jonathandgjones #PlantPathIGI
— kelsey wood (@klsywd) January 25, 2017
Marc Ghislan (CIP) also talked about Rpi-vnt1 for potato late blight resistance, this time in the context of field trials in Africa, where it was highly successful in preventing infection by Phytophthora infestans. He hopes that it will be available to grow soon for small-holder farms in Africa, where there is a huge yield gap due to disease.
Huge yield gap for potato for small holder farms – lose almost 1/2 yield to plant pathogens -Marc Ghislain @Cipotato #PlantPathIGI pic.twitter.com/UsqL5SKtM4
— kelsey wood (@klsywd) January 25, 2017
Cassava was the crop discussed in the next two talks, with the primary focus on the difficult to control viruses that infect them. Jim Carrington (Danforth Center) talked about how breeding for cassava mosaic virus resistance has accidentally increased susceptibility for cassava brown streak (which is in a different family of viruses). Linda Hanley-Bowdoin (North Carolina State) also talked about cassava mosaic virus and how complicated it can be to fight these viruses. Not content with simply mutating their genome and overcoming resistance like most pathogens, these viruses have also seemed to have evolved independently replicating sequences that enhance susceptibility to disease!
Thanks @klsywd for tweets from #PlantPathIGI
Report of Sequences Enhancing Geminivirus Susceptibility is published https://t.co/9chxyvDrUw— Steen Hoyer (@jshoyer) January 26, 2017
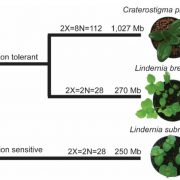
Doug Cook (UC Davis) presented impressive work on using genomics to understand variation in wild crop progenitors and their microbiomes. Leena Tripathi (International Institute of Tropical Agriculture) presented on several very effective strategies to control bacterial wilt on banana, which is a staple crop in Uganda, using genetic engineering.
Leena Tripathi grossing everyone out with photos of banana infected with xanthomonas wilt #PlantPathIGI #yuck pic.twitter.com/L6XFUhQ08h
— kelsey wood (@klsywd) January 26, 2017
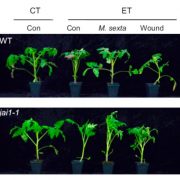
And finally, to end the symposium, Brian Staskawicz (UC Berkeley) gave a nice talk demonstrating how genome editing will be extremely useful for engineering plant resistance to pathogens. While many of the talks of the day featured resistance genes, Staskawicz showed promising results from experiments on tomato showing how knocking out susceptibility genes can lead to broad spectrum resistance to bacteria, fungi, and oomycetes all by targeting a single gene.
Promising results by knocking out a susceptibility gene using #CRISPR for broad spectrum disease resistance in ???? –@bstask #PlantPathIGI
— kelsey wood (@klsywd) January 26, 2017
All in all, it was a great symposium and I thank the organizers for inviting such a diverse array of speakers! I hope to see more symposia on genomics and genome editing and their applications to plant science come out of the Innovative Genomics Institute in the future!





Leave a Reply
Want to join the discussion?Feel free to contribute!