How a Mangrove tree can help to improve the salt tolerance of Arabidopsis and Rice
Affiliation: University of Melbourne
ORCiD: 0000-0001-5092-6168
email: [email protected]
Mangrove trees live and thrive in intertidal zones, where they are regularly inundated with salt water. To survive such harsh environmental conditions, they have evolved several features to improve their salt tolerance. Avicennia officinalis, which has increasingly been used in the past couple of years to study salinity tolerance, is a salt secretor mangrove tree with specialized salt glands in aboveground tissues to secrete excess salt. A. officinalis also has evolved enhanced hydrophobic barrier depositions in the endodermis of their roots. These barriers, namely the Casparian strips and suberin lamellae (SL), can reduce salt uptake from the environment into the vasculature by up to 95% (Tan et al., 2013; Krishnamurthy et al., 2017).
In a previous study, Krishnamurthy et al. (2017) identified several members of the Cytochrome P450 (CYP) gene family that are upregulated in the root transcriptome of A. officinalis in response to salt treatments. Out of those, AoCYP94B1 is among the family members that showed the strongest upregulation (Krishnamurthy et al., 2017). CYP94 subfamily members are fatty acid ω-hydroxylases, and AtCYP94B1 has been implicated in stress-induced jasmonic acid biosynthesis (Koo et al., 2014; Bruckhoff et al., 2016). However ω-hydroxylation is also required for suberin biosynthesis, leading Krishnamurthy et al. to investigate a possible role for AoCYP94B1 in suberin deposition to enhance barrier formation in the root.
In this issue of Plant Physiology, Krishnamurthy et al. (2020) functionally characterized CYP94B1 from A. officinalis, as well as its ortholog from Arabidopsis thaliana, and describe its conserved role in root suberin deposition for enhanced salt tolerance.
Having identified AoCYP94B1 in a transcriptomics screen, Krishnamurthy et al. analyzed the expression pattern of AoCYP94B1 and AtCYP94B1. Both orthologs are expressed in all tissues tested, with higher expression in the above-ground organs. Upon salt treatment, however, expression is strongly and rapidly induced in the root. As no transformation protocols are available for A. officinalis, the authors checked the expression and localization of the GFP-tagged Arabidopsis ortholog, and found that AtCYP94B1-GFP is expressed in the vasculature and endodermis of A. thaliana roots. In response to salt treatment the protein further accumulates at the plasma membrane of the endodermis, the location of apoplastic barrier formation.
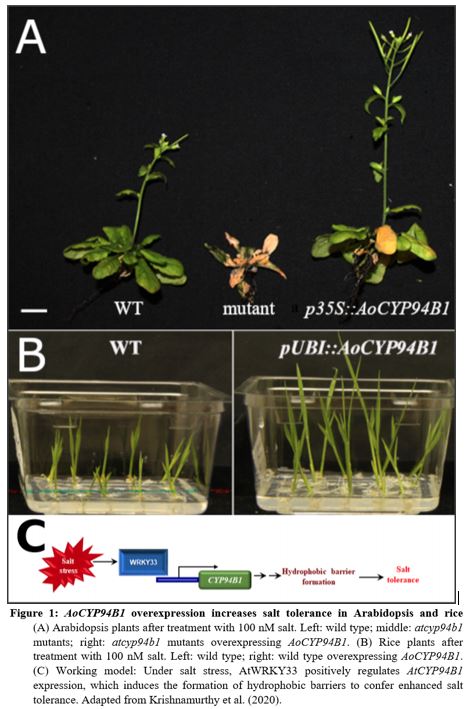 The authors analyzed Arabidopsis cyp94b1 mutants for their salt tolerance, and found that these mutants are indeed more sensitive to salt, while overexpression of either the AoCYP94B1 or AtCYP94B1 ortholog not only restores, but increases their tolerance (Fig. 1A). Similarly, overexpression of AoCYP94B1 in wild type rice also was sufficient to enhance the salt tolerance of this crop plant (Fig. 1B). Such overexpressing Arabidopsis lines accumulated less Na+ ions in leaves compared to the mutant, while root Na+ was unchanged, further indicating a role for AoCYP94B1 in formation of an apoplastic barrier that limits salt uptake into the root vasculature for subsequent transport to the above-ground organs. To confirm these observations, the authors screened for suberin monomers using mass spectrometry and found that the atcyp94b1 mutant had decreased amounts of root suberin monomers. Accordingly, staining with Nile Red showed decreased suberin in the endodermis of Arabidopsis mutant roots, and enhanced uptake of fluorescein diacetate (FDA) into the endodermis and pericycle. In rice (Oryza sativa), while mutants were not available, overexpression of AoCYP94B1 also resulted in stronger Nile Red staining of the endodermis, indicating enhanced suberin deposition.
The authors analyzed Arabidopsis cyp94b1 mutants for their salt tolerance, and found that these mutants are indeed more sensitive to salt, while overexpression of either the AoCYP94B1 or AtCYP94B1 ortholog not only restores, but increases their tolerance (Fig. 1A). Similarly, overexpression of AoCYP94B1 in wild type rice also was sufficient to enhance the salt tolerance of this crop plant (Fig. 1B). Such overexpressing Arabidopsis lines accumulated less Na+ ions in leaves compared to the mutant, while root Na+ was unchanged, further indicating a role for AoCYP94B1 in formation of an apoplastic barrier that limits salt uptake into the root vasculature for subsequent transport to the above-ground organs. To confirm these observations, the authors screened for suberin monomers using mass spectrometry and found that the atcyp94b1 mutant had decreased amounts of root suberin monomers. Accordingly, staining with Nile Red showed decreased suberin in the endodermis of Arabidopsis mutant roots, and enhanced uptake of fluorescein diacetate (FDA) into the endodermis and pericycle. In rice (Oryza sativa), while mutants were not available, overexpression of AoCYP94B1 also resulted in stronger Nile Red staining of the endodermis, indicating enhanced suberin deposition.
To gain insight into the transcriptional regulation of AoCYP94B1, the authors then screened the putative regulatory sequences of AoCYP94B1 for known transcription factor binding sites. Among the motifs identified, they found several potential WRKY binding sites, and the authors’ previous transcriptomic data showed an upregulation of several WRKYs after salt stress, most notably WRKY33 (Krishnamurthy et al., 2017). WRKY33 has previously been implicated as a key regulator of biotic and abiotic stresses in Arabidopsis, and its ortholog in the halophyte Eutrema salsugineum has been found to be important for the plant’s enhanced salt tolerance (Jiang and Deyholos, 2009; Birkenbihl et al., 2012; Mucha et al., 2015). Accordingly, the authors focused on WRKY33 as a potential transcriptional regulator of AtCYP94B1. Indeed, they found that atwrky33 mutants show reduced suberin deposition in their roots, are more sensitive to salt stress, and express AtCYP94B1 at lower levels than the wild type. Overexpression of AtCYP94B1 in wrky33 mutants restored these phenotypes, indicating that the role of CYP94B1 in enhancing salt tolerance by increasing suberin deposition is indeed dependent on positive transcriptional regulation by WRKY33. Using chromatin immunoprecipitation-quantitative PCR and yeast one-hybrid assays they then also confirmed that the AtCYP94B1 regulatory sequences are bound by AtWRKY33.
The work described in this article presents CYP94B1 as a central regulator of salt tolerance in a WRKY33-dependent pathway that furthermore seems to be conserved among several, distantly related plant species, from a mangrove tree to Arabidopsis to rice (Fig. 1C). Previous work has implicated WRKY33 as a key transcriptional regulator not just for salt tolerance, but for a whole range of biotic and abiotic stresses, most notably resistance towards the fungal pathogen Botrytis cinerea (Jiang and Deyholos, 2009; Birkenbihl et al., 2012). In this regard, WRKY33 has been shown to act by regulating phytohormones such as jasmonic acid and salicylic acid. Since CYP94B1 has also been implicated to function in stress-induced jasmonic acid biosynthesis, and suberin-deposition also protects from pathogen invasion, it could be interesting to further investigate how WRKY33 and CYP94B1 integrate different pathways, in response to a variety of stresses.
LITERATURE CITED
Birkenbihl RP, Diezel C, Somssich IE (2012) Arabidopsis WRKY33 is a key transcriptional regulator of hormonal and metabolic responses toward Botrytis cinerea infection. Plant Physiol 159: 266–85
Bruckhoff V, Haroth S, Feussner K, König S, Brodhun F, Feussner I (2016) Functional Characterization of CYP94-Genes and Identification of a Novel Jasmonate Catabolite in Flowers. PLoS One 11: e0159875
Jiang Y, Deyholos MK (2009) Functional characterization of Arabidopsis NaCl-inducible WRKY25 and WRKY33 transcription factors in abiotic stresses. Plant Mol Biol 69: 91–105
Koo AJ, Thireault C, Zemelis S, Poudel AN, Zhang T, Kitaoka N, Brandizzi F, Matsuura H, Howe GA (2014) Endoplasmic reticulum-associated inactivation of the hormone jasmonoyl-L-isoleucine by multiple members of the cytochrome P450 94 family in Arabidopsis. J Biol Chem 289: 29728–38
Krishnamurthy P, Mohanty B, Wijaya E, Lee D-Y, Lim T-M, Lin Q, Xu J, Loh C-S, Kumar PP (2017) Transcriptomics analysis of salt stress tolerance in the roots of the mangrove Avicennia officinalis. Sci Rep 7: 10031
Krishnamurthy P, Vishal B, Ho WJ, Lok FCJ, Lee FSM, Kumar PP (2020) Regulation of a cytochrome P450 gene CYP94B1 by WRKY33 transcription factor controls apoplastic barrier formation in roots to confer salt tolerance. Plant Physiol 7303: pp.01054.2020 https://doi.org/10.1104/pp.20.01054
Mucha S, Walther D, Müller TM, Hincha DK, Glawischnig E (2015) Substantial reprogramming of the Eutrema salsugineum (Thellungiella salsuginea) transcriptome in response to UV and silver nitrate challenge. BMC Plant Biol 15: 137
Tan W-K, Lin Q, Lim T-M, Kumar P, Loh C-S (2013) Dynamic secretion changes in the salt glands of the mangrove tree species Avicennia officinalis in response to a changing saline environment. Plant Cell Environ 36: 1410–22