High resolution mapping of RphMBR1012 conferring resistance to Puccinia hordei in barley (Front Plant Sci)
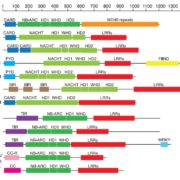
 Barley leaf rust (Puccinia hordei) is a biotrophic fungus, in which mutations cause newly virulent races (pathothypes) with severe effects on cultivars. Marker-assisted Selection for Rph (Resistance to P. hordei) genes has been studied to provide sustainable control of this disease. Fazlikhani et al. developed molecular markers for the construction of a high resolution map for the RphMBR1012 locus on recombinant inbred lines (RILs), derived from the cross of MBR1012 (resistant) x Scarlett (susceptible) lines. Fifty-six markers were identified/developed using New Illumina SNP genotyping analysis and Genotyping-by-Sequencing (GBS). Marker saturation of the pathogen resistance associated locus and its physical mapping were performed thanks to the barley reference Zipper genome sequence. The authors increased the genetic resolution by the conversion into KASP markers (Kompetitive Allele Specific PCR), which narrowed down the RphMBR1012 target interval to 0.44 Mb between the markers QBS127 and QBS98. In this interval, five candidate R genes were identified and analyzed by allele specific re-sequencing. The markers GBS546 and GBS626 were catalogued as the best to RphMBR1012 detection in barley. (Summary by Ana Valladares). Front Plant Sci. 10.3389/fpls.2019.00640.
Barley leaf rust (Puccinia hordei) is a biotrophic fungus, in which mutations cause newly virulent races (pathothypes) with severe effects on cultivars. Marker-assisted Selection for Rph (Resistance to P. hordei) genes has been studied to provide sustainable control of this disease. Fazlikhani et al. developed molecular markers for the construction of a high resolution map for the RphMBR1012 locus on recombinant inbred lines (RILs), derived from the cross of MBR1012 (resistant) x Scarlett (susceptible) lines. Fifty-six markers were identified/developed using New Illumina SNP genotyping analysis and Genotyping-by-Sequencing (GBS). Marker saturation of the pathogen resistance associated locus and its physical mapping were performed thanks to the barley reference Zipper genome sequence. The authors increased the genetic resolution by the conversion into KASP markers (Kompetitive Allele Specific PCR), which narrowed down the RphMBR1012 target interval to 0.44 Mb between the markers QBS127 and QBS98. In this interval, five candidate R genes were identified and analyzed by allele specific re-sequencing. The markers GBS546 and GBS626 were catalogued as the best to RphMBR1012 detection in barley. (Summary by Ana Valladares). Front Plant Sci. 10.3389/fpls.2019.00640.