An Alternative Route for Astaxanthin Biosynthesis in Green Algae
Tianhu Sun
ORCID ID: 0000-0002-2513-1387
Plant Breeding and Genetics Section, School of Integrative Plant Science, Cornell University, Ithaca, New York 14853
Astaxanthin is the reddish carotenoid pigment that gives color to shrimp, salmon, and flamingo. However, these animals don’t synthesize astaxanthin by themselves. Instead, microalgae are the initial producers of astaxanthin in nature. Because of its strong antioxidant activity and potential benefits for human health, astaxanthin has high commercial value in the nutraceutical, cosmetic, and feed industries. Haematococcus pluvialis (H. pluvialis) is a well-studied alga with the highest known level of astaxanthin accumulation and thus is widely used as an industrial source for natural astaxanthin production. In recent years, Chromochloris zofingiensis (C. zofingiensis) has emerged as a promising producer because of its ability to synthesize astaxanthin and its better growth robustness than H. pluvialis. Compared to H. pluvialis, relatively few genes have been characterized from C. zofingiensis. With the availability of the whole-genome sequence, the identification of structural genes involved in carotenoid biosynthesis in C. zofingiensis become feasible (Roth et al., 2017). However, the biosynthetic route of astaxanthin in C. zofingiensis remains ambiguous. In this issue of Plant Physiology, Zhang et al. (2020) elucidate the pathways for astaxanthin biosynthesis and accumulation in C. zofingiensis, which shed light on astaxanthin production in green algae.
As for other carotenoids, the backbone of astaxanthin is built from the five-carbon precursors isopentenyl diphosphate (IPP) and dimethylallyl pyrophosphate (DMAPP). Two separate pathways, the cytosolic mevalonate (MVA) pathway and the chloroplastic methylerythritol phosphate (MEP) pathway, can provide these precursors (Rodriguez-Concepcion et al., 2018). Zhang et al. (2020) showed that the precursors of astaxanthin are derived from the MEP pathway rather than the MVA pathway: when inhibitors for each pathway were tested on C. zofingiensis, only MEP pathway inhibition affected astaxanthin accumulation. This is the same precursor supply pathway as used in H. pluvialis. Interestingly, the expression of key enzymes controlling the MEP pathway was not induced by nitrogen deprivation in C. zofingiensis like it is in H. pluvialis. It is possible that the biosynthesis of the smaller amounts of astaxanthin in C. zofingiensis is not limited by the precursor supply from the MEP pathway as in H. pluvialis, consistent with the lack of induction of the MEP biosynthetic genes.
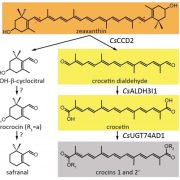
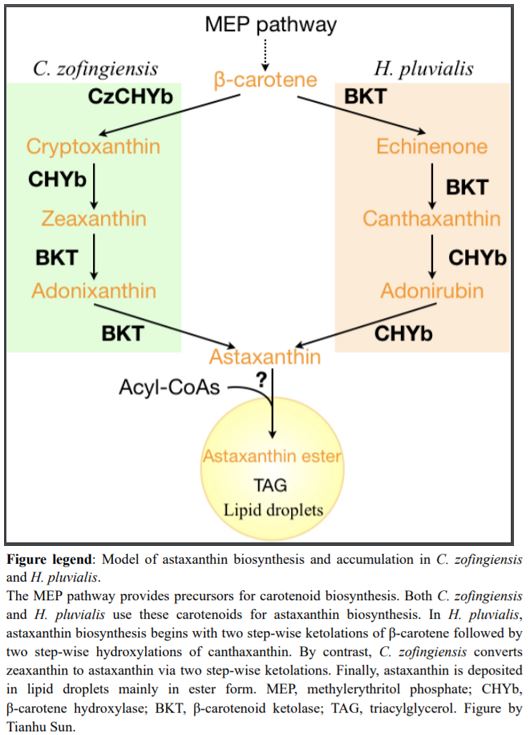 In H. pluvialis, astaxanthin biosynthesis begins with β-carotene followed by two ketolation steps catalyzed by β-carotene ketolase (BKT) to produce canthaxanthin, and then two hydroxylation steps of canthaxanthin catalyzed by β-carotene hydroxylase (CHYb) (Figure 1) (Schoefs et al., 2001). Ketolation is the key step for astaxanthin production, and land plants cannot produce astaxanthin because they lack BKT. Inhibition or mutation of BKT in C. zofingiensis abolished the production of all kinds of keto-carotenoids (Roth et al., 2017; Ye and Huang, 2020). Unexpectedly, zeaxanthin also accumulated, suggesting that C. zofingiensis may have an unidentified route to produce astaxanthin other than that used by H. pluvialis (Roth et al., 2017; Ye and Huang, 2020).
In H. pluvialis, astaxanthin biosynthesis begins with β-carotene followed by two ketolation steps catalyzed by β-carotene ketolase (BKT) to produce canthaxanthin, and then two hydroxylation steps of canthaxanthin catalyzed by β-carotene hydroxylase (CHYb) (Figure 1) (Schoefs et al., 2001). Ketolation is the key step for astaxanthin production, and land plants cannot produce astaxanthin because they lack BKT. Inhibition or mutation of BKT in C. zofingiensis abolished the production of all kinds of keto-carotenoids (Roth et al., 2017; Ye and Huang, 2020). Unexpectedly, zeaxanthin also accumulated, suggesting that C. zofingiensis may have an unidentified route to produce astaxanthin other than that used by H. pluvialis (Roth et al., 2017; Ye and Huang, 2020).
To investigate the contribution of BKT in astaxanthin production in C. zofingiensis, Zhang et al. (2020) started by detecting metabolic intermediates to reconstruct the astaxanthin biosynthetic pathway. Interestingly, carotenoid profiling in C. zofingiensis detected the β-carotene ketolation product canthaxanthin but not the further hydroxylation intermediate adonirubin, which in H. pluvialis is catalyzed by CHYb to generate astaxanthin. Subsequent in vitro enzymatic assays of BKT and CHYb from C. zofingiensis revealed that BKT can catalyze both β-carotene to canthaxanthin and zeaxanthin to astaxanthin, while CHYb can only catalyze β-carotene to zeaxanthin but not canthaxanthin to astaxanthin. Therefore, the astaxanthin biosynthesis in C. zofingiensis is proposed to begin with CHYb catalyzing β-carotene to zeaxanthin and then BKT catalyzing zeaxanthin to astaxanthin, which is different from H. pluvialis (Figure 1).
Where and in which form carotenoids are stored are equally important as their biosynthesis to final carotenoid content (Sun et al. 2018). Zhang et al. (2020) found that astaxanthin, mostly in ester form, accumulated in lipid droplets after stress induction in C. zofingiensis. Triacylglycerol (TAG) constitutes the hydrophobic core of lipid droplets. A good correlation between TAG content, astaxanthin accumulation, and TAG synthesis-related and carotenogenic gene expression was observed.
Whether fatty acid biosynthesis is a key factor for astaxanthin accumulation was further investigated. In H. pluvialis, inhibition of de novo fatty acid biosynthesis severely decreases astaxanthin biosynthesis (Chen et al., 2015). However, this is not the case in C. zofingiensis. Instead, Zhang et al. (2020) showed that inhibition of de novo fatty acid biosynthesis elevated astaxanthin production without affecting the expression of astaxanthin biosynthetic pathway genes. Furthermore, feeding of free fatty acid did not affect astaxanthin biosynthesis. Thus, Zhang et al. (2020) conclude that the storage of astaxanthin in lipid droplets is coordinated with TAG accumulation but independent from the de novo fatty acid biosynthesis in C. zofingiensis.
In summary, this study proposes that the biosynthetic pathway for astaxanthin in C. zofingiensis is different from the well-studied pathway in H. pluvialis and also dissects the deposition of astaxanthin in C. zofingiensis. Most importantly, Zhang et al. (2020) reveal that C. zofingiensis BKT can use zeaxanthin as a substrate to generate astaxanthin, bypassing the bottleneck in the hydroxylation of canthaxanthin by CHYb, providing us fundamental knowledge to improve and manipulate astaxanthin biosynthesis in the future.
These findings also raise some additional questions. First, what are the key amino acid residues for the enzymatic activities of CHYb and BKT? This knowledge can lead to the improvement of enzyme specificity and efficiency to generate better algal strains for astaxanthin production. Since plants produce abundant zeaxanthin, a turbo-charged BKT enzyme that can convert zeaxanthin to astaxanthin could generate astaxanthin-rich crops via a single gene modification. Moreover, it remains to be elucidated why astaxanthin accumulation is independent of de novo fatty acid biosynthesis and how to increase the storage capacity for astaxanthin in C. zofingiensis.
LITERATURE CITED
Chen G, Wang B, Han D, Sommerfeld M, Lu Y, Chen F, Hu Q (2015) Molecular mechanisms of the coordination between astaxanthin and fatty acid biosynthesis in Haematococcus pluvialis (Chlorophyceae). Plant J 81: 95-107
Rodriguez-Concepcion M, Avalos J, Bonet ML, Boronat A, Gomez-Gomez L, Hornero-Mendez D, Limon MC, Melendez-Martinez AJ, Olmedilla-Alonso B, Palou A, Ribot J, Rodrigo MJ, Zacarias L, Zhu C (2018) A global perspective on carotenoids: Metabolism, biotechnology, and benefits for nutrition and health. Prog Lipid Res 70: 62-93
Roth MS, Cokus SJ, Gallaher SD, Walter A, Lopez D, Erickson E, Endelman B, Westcott D, Larabell CA, Merchant SS, Pellegrini M, Niyogi KK (2017) Chromosome-level genome assembly and transcriptome of the green alga Chromochloris zofingiensis illuminates astaxanthin production. Proc Natl Aca Sci USA 114: 4296-4305
Schoefs B, Rmiki N-E, Rachadi J, Lemoine Y (2001) Astaxanthin accumulation in Haematococcus requires a cytochrome P450 hydroxylase and an active synthesis of fatty acids. FEBS Lett 500: 125-128
Sun T, Yuan H, Cao H, Yazdani M, Tadmor Y, Li L (2018) Carotenoid metabolism in plants: the role of plastids. Mol Plant 11: 58-74
Ye Y, Huang J-C (2020) Defining the biosynthesis of ketocarotenoids in Chromochloris zofingiensis. Plant Diversity 42: 61-66