What We’re Reading: June 30th
 This edition of What We’re Reading is guest edited by Mather A Khan, a Postdoc Fellow in the Mendoza Lab at Bond Life Sciences Center, University of Missouri, Columbia, USA. He is very much concerned about hunger, poverty, and global food security. His research is always been motivated by these factors.
This edition of What We’re Reading is guest edited by Mather A Khan, a Postdoc Fellow in the Mendoza Lab at Bond Life Sciences Center, University of Missouri, Columbia, USA. He is very much concerned about hunger, poverty, and global food security. His research is always been motivated by these factors.
“Plant Science research always addresses the very basic necessity of life. Understanding the fundamentals of growth and development of plants is of utmost importance in order to meet the global demands for food.” Mather is working to identify and characterize the molecular players involved in metal homeostasis and to unravel the molecular mechanisms behind it.
Review: Node-controlled allocation of mineral elements ($)
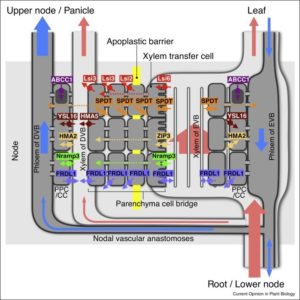 Essential metals and non-essential metals and metalloids enter the plant system from the soil through roots and are distributed to different parts of the plants by xylem and phloem, primarily through transpiration flow. In gramineous plants like rice and barley, nodes play a very important role in nutrient allocation. This review, by Yamaji and Ma, is focused on nodes and their role in mineral nutrient transport and allocation. The authors discuss node structures, anatomical features of node vasculature, and transporters involved in loading and unloading. They also discuss the regulation of node-localized transporters. The authors highlight that “manipulation of node-based transporters will provide a novel breeding target to improve nutrient use efficiency, productivity, nutritional value and safety in cereal crops.” Curr. Opin. Plant Biol. 10.1016/j.pbi.2017.05.002
Essential metals and non-essential metals and metalloids enter the plant system from the soil through roots and are distributed to different parts of the plants by xylem and phloem, primarily through transpiration flow. In gramineous plants like rice and barley, nodes play a very important role in nutrient allocation. This review, by Yamaji and Ma, is focused on nodes and their role in mineral nutrient transport and allocation. The authors discuss node structures, anatomical features of node vasculature, and transporters involved in loading and unloading. They also discuss the regulation of node-localized transporters. The authors highlight that “manipulation of node-based transporters will provide a novel breeding target to improve nutrient use efficiency, productivity, nutritional value and safety in cereal crops.” Curr. Opin. Plant Biol. 10.1016/j.pbi.2017.05.002
Review: New tool for plant genome editing ($)
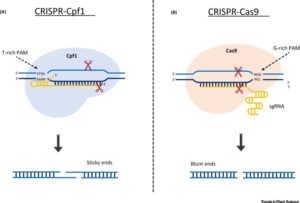 The genome editing tool CRISPR-Cas9 has been successfully applied in many organisms including crop plants such as rice, wheat, maize, tomato, potato, tobacco, cotton, and soybean. This review is on a different CRISPR-based gene editing system, CRSPR-Cpf1. Cpf1 is an endonuclease found in Prevotella and Francisella bacteria. The authors presents a comparison between CRISPR-Cas9 and CRISPR-Cpf1. They also discuss several recent studies on CRISPR-Cpf1 showing greater efficiency over the CRISPR-Cas9 system. Trends Plant Sci. 10.1016/j.tplants.2017.05.001
The genome editing tool CRISPR-Cas9 has been successfully applied in many organisms including crop plants such as rice, wheat, maize, tomato, potato, tobacco, cotton, and soybean. This review is on a different CRISPR-based gene editing system, CRSPR-Cpf1. Cpf1 is an endonuclease found in Prevotella and Francisella bacteria. The authors presents a comparison between CRISPR-Cas9 and CRISPR-Cpf1. They also discuss several recent studies on CRISPR-Cpf1 showing greater efficiency over the CRISPR-Cas9 system. Trends Plant Sci. 10.1016/j.tplants.2017.05.001
Host plants synthesize fatty acids for symbiotic fungi ($)
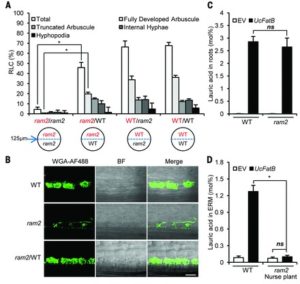 The symbiotic association between plants and arbuscular mycorrhizal fungi is well known. The fungi facilitate mineral nutrient acquisition from the soil, while the plants support fungi by providing carbon sources in the form of carbohydrates. Jiang et al. report that in this symbiotic relationship, in addition to providing sugars, the plants also supplies fatty acids to its fungi to sustain mycorrhizal colonization. Science 10.1126/science.aam9970
The symbiotic association between plants and arbuscular mycorrhizal fungi is well known. The fungi facilitate mineral nutrient acquisition from the soil, while the plants support fungi by providing carbon sources in the form of carbohydrates. Jiang et al. report that in this symbiotic relationship, in addition to providing sugars, the plants also supplies fatty acids to its fungi to sustain mycorrhizal colonization. Science 10.1126/science.aam9970
LjMOT1, a high-affinity molybdate transporter from Lotus japonicas ($)
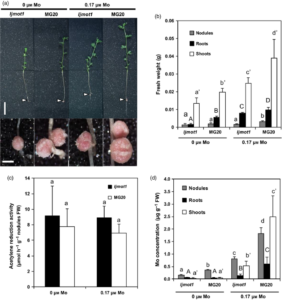 Molybdenum (Mo) is an essential micronutrient found in the soil that is required for growth and development by most biological organisms including plants and animals. It is biologically inactive until bound with pterin to form Mo-cofactor (Moco). In this article, Duan et al. report a high-affinity Mo transporter (LjMOT1) from L. japonicas. The transport activity of LjMOT1 was characterized by expressing it in yeast and in planta. Yeast cells expressing LjMOT1 showed a 100-fold increase in Mo concentrations, indicating that the transporter has Mo uptake activity. Plant J.
Molybdenum (Mo) is an essential micronutrient found in the soil that is required for growth and development by most biological organisms including plants and animals. It is biologically inactive until bound with pterin to form Mo-cofactor (Moco). In this article, Duan et al. report a high-affinity Mo transporter (LjMOT1) from L. japonicas. The transport activity of LjMOT1 was characterized by expressing it in yeast and in planta. Yeast cells expressing LjMOT1 showed a 100-fold increase in Mo concentrations, indicating that the transporter has Mo uptake activity. Plant J.
Imaging and metabolite profiling in maize seed germination
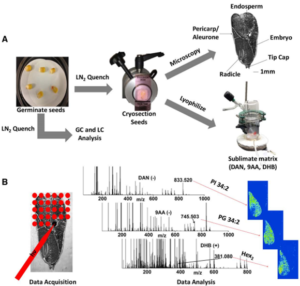 Seed germination is a complex developmental process. Decoding it is of practical importance to improve agriculture. During seed germination, massive changes of metabolite levels occur. In this work, a combination of GC-MS, LC-fluorescence, and mass spectrometry imaging (MSI) approaches were used for quantitative metabolite profiling and to visualize metabolite distributions during germination of seeds of two different maize inbred lines, M017 and B73, at different times post-imbibition. Feenstra et al. conclude “this study demonstrates the synergy that can be gained by combining these strategies to reveal new insights into complex biological processes that integrate capabilities of different cellular compartments.” Plant Physiol.
Seed germination is a complex developmental process. Decoding it is of practical importance to improve agriculture. During seed germination, massive changes of metabolite levels occur. In this work, a combination of GC-MS, LC-fluorescence, and mass spectrometry imaging (MSI) approaches were used for quantitative metabolite profiling and to visualize metabolite distributions during germination of seeds of two different maize inbred lines, M017 and B73, at different times post-imbibition. Feenstra et al. conclude “this study demonstrates the synergy that can be gained by combining these strategies to reveal new insights into complex biological processes that integrate capabilities of different cellular compartments.” Plant Physiol.
HMA3 confers cadmium hypertolerance to Sedum lumbizincicola ($)
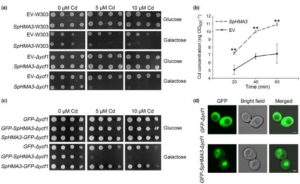 Cadmium (Cd) is a non-essential heavy metal that is toxic to most organisms. Some plants, called hyperaccumulators, actively uptake large amount of toxic heavy metals including Cd. Hyperaccumulator plants have a very efficient detoxification system to protect against the lethal effect of toxic metals and are excellent candidates for phytoremediation (the removal of toxins from soil by plants). The mechanism of detoxification including of Cd is poorly understood. Liu et al. showed that SpHMA3, localized to the tonoplast, plays a very important role in Cd detoxification in young leaves and stems by sequestering Cd into vacuoles. Using yeast as a heterologous expression system, they show that SpHMA3 transports Cd but not Zn, Pb, Co, or Mn. RNAi knockdown lines of S. plumbizincicola are sensitive to Cd but not to Zn, indicating that SpHMA3 confers Cd hypertolerance to S. plumbizincicola. New Phytol.
Cadmium (Cd) is a non-essential heavy metal that is toxic to most organisms. Some plants, called hyperaccumulators, actively uptake large amount of toxic heavy metals including Cd. Hyperaccumulator plants have a very efficient detoxification system to protect against the lethal effect of toxic metals and are excellent candidates for phytoremediation (the removal of toxins from soil by plants). The mechanism of detoxification including of Cd is poorly understood. Liu et al. showed that SpHMA3, localized to the tonoplast, plays a very important role in Cd detoxification in young leaves and stems by sequestering Cd into vacuoles. Using yeast as a heterologous expression system, they show that SpHMA3 transports Cd but not Zn, Pb, Co, or Mn. RNAi knockdown lines of S. plumbizincicola are sensitive to Cd but not to Zn, indicating that SpHMA3 confers Cd hypertolerance to S. plumbizincicola. New Phytol.
Short-term high temperature response in Pinus radiata ($)
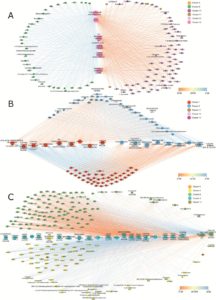 Plants have the ability to perceive and respond under changing environmental conditions including a rise in temperature. In the current context of climate change and global warming, understanding the physiological and molecular mechanism of heat response is very important. This study is on the high-temperature response in Pinus radiata, the most widely planted pine species. Escandon et al. studied the short-term heat response by exploring the metabolome, proteome and targeted transcriptome in P. radiata. The authors report the identification of 901 protein groups and 2075 metabolites by using different ‘omics’ tools. J. Exp. Bot. 10.1093/jxb/erx198
Plants have the ability to perceive and respond under changing environmental conditions including a rise in temperature. In the current context of climate change and global warming, understanding the physiological and molecular mechanism of heat response is very important. This study is on the high-temperature response in Pinus radiata, the most widely planted pine species. Escandon et al. studied the short-term heat response by exploring the metabolome, proteome and targeted transcriptome in P. radiata. The authors report the identification of 901 protein groups and 2075 metabolites by using different ‘omics’ tools. J. Exp. Bot. 10.1093/jxb/erx198
Genome-wide mutant library in rice
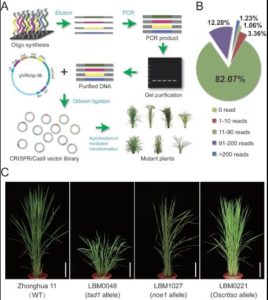 Rice is a staple food for nearly half of the world’s people. It is also a powerful model system for studying monocots due its relatively small genome size and the availability of a complete genome sequence. The next challenge is to functionally characterize each gene, which is very important for functional genomics and for improving traits in rice. Meng et al. used the CRISPR-Cas9 system to construct a genome-wide mutant library in rice. To generate the large scale mutant library, the authors chose the first two identified sgRNA target sites in each candidate gene. They selected 12,802 genes highly expressed in rice shoot basal tissue and 25,604 corresponding sgRNAs. Overall, this is a very useful resource for the researchers working on rice. Mol. Plant. 10.1016/j.molp.2017.06.006
Rice is a staple food for nearly half of the world’s people. It is also a powerful model system for studying monocots due its relatively small genome size and the availability of a complete genome sequence. The next challenge is to functionally characterize each gene, which is very important for functional genomics and for improving traits in rice. Meng et al. used the CRISPR-Cas9 system to construct a genome-wide mutant library in rice. To generate the large scale mutant library, the authors chose the first two identified sgRNA target sites in each candidate gene. They selected 12,802 genes highly expressed in rice shoot basal tissue and 25,604 corresponding sgRNAs. Overall, this is a very useful resource for the researchers working on rice. Mol. Plant. 10.1016/j.molp.2017.06.006



Leave a Reply
Want to join the discussion?Feel free to contribute!