Turnover of Tonoplast Proteins
By
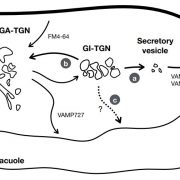
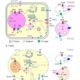
Our knowledge of vacuole biogenesis and the transport of proteins to the vacuole has advanced consistently over the last 30 years. In meristematic cells, the tonoplast appears to develop directly out of the endoplasmic reticulum (Viotti et al., 2013). Once it is established as a functioning organelle, tonoplast proteins reach the vacuole in one of two main ways. A bulk flow process for tonoplast protein delivery originates at the trans-Golgi network (TGN) and proceeds through maturation into a multivesicular endosome that ultimately fuses with the vacuole (Brillada and Rojas-Pierce, 2017). Alternatively, vesicle transport from the TGN targets tonoplast proteins to this endosomal compartment. Studies using adaptor protein complex (AP) mutants indicate that the transport vectors in this latter case may be clathrin-coated vesicles. There is also some evidence to suggest that some of the TGN-derived vesicles (AP-4-positive, but lacking clathrin) might bypass the endosomes (Rosquete et al., 2018). In accordance with this notion, it has been recently demonstrated that two different tethering complexes (CORVET and HOPS) exist for fusion with the tonoplast in Arabidopsis (Arabidopsis thaliana; Takemoto et al., 2018). Thus, transport to the vacuole would appear to be more complicated than was previously thought.
Plant cells maintain vacuoles for long timespans, and they continuously receive new membrane in the form of fusing vesicles or endosomes. Since vacuoles cannot enlarge indefinitely, they must somehow compensate by removing excess membrane proteins and lipids. This may occur by way of microautophagy, sometimes leading to “bulb” formation (Han et al., 2015; Maîtrejean and Vitale, 2012). Preliminary evidence indicating that bulb formation might be a selective rather than a bulk degradative process has been published (Saito et al., 2002). A possible selective removal of proteins from the tonoplast is also indirectly implied in the recent publication of Feeney et al. (2018), which documents the delivery of typical tonoplast proteins from the storage vacuole to lytic vacuoles during vacuole remodeling in Arabidopsis leaves. Exactly how such a transfer could occur is unclear, although vesicle-mediated retrograde transport might be inferred from the presence of a large set of endomembrane trafficking components in the vacuolar proteome, including small GTPases of the Arf and Rab7 classes (Carter et al., 2004).
Remarkable new insights into how membrane proteins may be removed from the tonoplast have recently been provided by the group of Scott Emr working on the yeast vacuole. In one publication, Suzuki and Emr (2018) investigated the retrieval of the type I membrane protein Atg27 from the yeast vacuole membrane. Normally, this protein functions together with Atg9 in the formation of microvesicles that are required for autophagosome formation. However, by virtue of a Tyr-based motif in its cytoplasmic C-tail, it interacts with the AP3 adaptor complex and also gets transported directly from the Golgi to the vacuole. By generating yeast mutants with impaired retromer (vps35Δ) or sorting nexin (snx4Δ) function, Suzuki and Emr were able to show that retrograde transport of Atg27 from the vacuole to the Golgi occurs in two steps: First, Atg27 is incorporated into a vacuole-to-endosome transport vesicle formed by the attachment of dimers of Snx4 and Snx41/42 via their PHOX homology (PX) and Bin/Amphiphysin/Rvs (BAR) domains on the vacuole membrane; second, a classical retromer-mediated transport of Atg27 from the endosome to the Golgi occurs. A direct interaction between Atg27 and Snx4 was demonstrated by coimmunoprecipitation, with R224, S227, F231, and I236 being determined as the responsible residues in the C-tail of Atg27. Thus, as with the retrograde transport of mannosyl 6-phosphate receptors from maturing endosomes in mammalian cells (for a discussion, see Robinson, 2018), it is once again the sorting nexins that have been demonstrated to be the crucial coat proteins for vesicle transport.
In another landmark article, Zhu et al. (2017) followed the degradation of the Lys transporter Ypq1. This degradation is initiated upon Lys withdrawal from the growth medium and causes a type E3 ubiquitin ligase, RSP5, to be recruited to the surface of the yeast vacuole. After being ubiquitinated, Ypq1 is sorted into a microvesicle and released into the vacuole lumen. Based on multiple genetic approaches, Zhu et al. (2017) concluded that the ESCRT (endosomal sorting complexes required for transport) complexes required for this internalization process are located directly on the vacuole membrane, in addition to those on endosomes as previously described (Henne et al., 2011).
At present, it is unclear how evolutionary conserved the Snx4 retrieval and vacuole-based ESCRT internalization mechanisms are, but we feel that they are unlikely to be restricted to yeast. Might they operate in plants? Unfortunately, Atg27 (YJL178C) has no homologs in the green lineage (Michaeli et al., 2016), while homologs of Snx41p (YDR425W) and Snx42p (YDL113C) may exist in green algae but not in plants. However, the situation with Snx4p (YJL036W) is different. BLAST searches reveal that the known BAR-domain SNX proteins of plants are distantly related to both Vps5p (YOR069W) and Snx4p, and therefore might have the capacity to perform recycling of tonoplast proteins. Two considerations must be taken into account. On the one hand, SNX1 (At5g06140) and SNX2 (At5g58440 and At5g07120) proteins mainly localize to the TGN (Stierhof et al., 2013); thus, their involvement at the tonoplast is not directly evident. On the other hand, one cannot exclude a potential short-lived SNX1/SNX2 tonoplast association and a rapid vesicle release that remains unnoticed in localization studies. Notably, Snx4 is also not obviously localized to the yeast vacuolar membrane (Suzuki and Emr, 2018). In support of a potential SNX1/SNX2 involvement is the fact that PI3P, the lipid targeted by the PX domain of SNX proteins, has been detected at the tonoplast (Simon et al., 2014). Moreover, SNX1 and SNX2 have been shown form homo- and heterodimers in vivo. Therefore, the absence of plant Snx41/42 homologs and the fact that the three Arabidopsis non-BAR-domain SNXs might not participate in membrane tubulation (for review, see Heucken and Ivanov, 2018) suggest that there might be no requirement for additional partners as in the case of Snx4p. However, before this can be addressed experimentally, we will first need to identify retrograde cargo proteins in plants.
Except for ESCRT complex 0, which is absent in plants, all of the other ESCRT complexes have been identified and functionally characterized in relation to the vacuolar degradation of ubiquitinated plant plasma membrane proteins (Nagel et al., 2017). However, and as in other eukaryotes, the ESCRT complexes in plants are located on endosomal membranes. To our knowledge, there is no publication showing the presence of ESCRT proteins at the tonoplast or that ubiquitination of specific proteins occurs at this location. For the moment, it would seem that this particular mechanism for tonoplast protein turnover is peculiar to yeast.
REFERENCES
Brillada C, Rojas-Pierce M (2017) Vacuolar traffivcking and biogenesis: a maturation in the field. Curr Opin Plant Biol 40: 77–81
Carter C, Pan S, Zouhar J, Avila EL, Girke T, Raikhel NV (2004) The vegetative vacuole proteome of Arabidopsis thaliana reveals predicted and unexpected proteins. Plant Cell 16: 3285–3303
Feeney M, Kittelmann M, Menassa R, Hawes C, Frigerio L (2018) Protein storage vacuoles originate by remodelling of preexisting vacuoles in Arabidopsis thaliana. Plant Physiol 177: 241–254
Han SW, Alonso JM, Rojas-Pierce M (2015) Regulator of bulb biogenesis 1 (RBB1) is involved in vacuole bulb formation in Arabidopsis. PLoS One 10: e0125621
Henne WM, Buchkovich NJ, Emr SD (2011) The ESCRT Pathway. Dev Cell 21: 77–91
Heucken N, Ivanov R (2018) The retromer, sorting nexins and the plant endomembrane protein trafficking. J Cell Sci 131: jcs203695
Michaeli S, Galili G, Genschik P, Fernie AR, Avin-Wittenberg T (2016) Autophagy in plants: What’s new on the menu? Trends Plant Sci 21: 134–144
Maîtrejean M, Vitale A (2012) How are tonoplast proteins degraded? Plant Signal Behav 6: 1809–1812
Nagel M-K, Kalinowska K, Vogel K, Reynolds GD, Wu Z, Anzenberger F, Ichikawa M, Tsutsumi C, Sato MH, Kuster B, Bednarek SY, Isono E (2017) Arabidopsis SH3P2 is an ubiquitin-binding protein that functions together with ESCRT-I and the deubiquitylating enzyme AMSH3. Proc Natl Acad Sci USA 114: E7197–E7204
Robinson DG (2018) Retromer and VSR recycling: A Red Herring? Plant Physiol 176: 483–484
Rosquete MR, Davis DJ, Drakakaki G (2018) The plant trans-Golgi network. Not just a matter of distinction. Plant Physiol 176: 187–198
Takemoto K, Ebine K, Askani JC, Krüger F, Gonzalez ZA, Ito E, Goh T, Schumacher K, Nakano A, Ueda T (2018) Distinct sets of tethering complexes, SNARE complexes, and Rab GTPases mediate membrane fusion at the vacuole in Arabidopsis. Proc Natl Acad Sci USA 115: E2457–E2466
Seito C, Ueda T, Aba H, Wada Y, Kuroiwa T, Hisada A, Furuya M, Nakano A (2002) A complex and mobile structure forms a distinct subregion within the continuous vacuolar membrane in young cotyledons of Artabidopsis. Plant J 29: 245–255
Simon ML, Platre MP, Assil S, van Wijk R, Chen WY, Chory J, Dreux M, Munnik T, Jaillais Y (2014) A multi-colour/ multi-affinity marker set to visualize phosphoinositide dynamics in Arabidopsis. Plant J 77: 322–337
Stierhof YD, Viotti C, Scheuring D, Sturm S, Robinson DG (2013) Sorting nexins 1 and 2a locate mainly to the TGN. Protoplasma 250: 235–240
Suzuki SW, Emr SD (2018) Membrane protein recycling from the vacuole/lysosome membrane. J Cell Biol pii: jcb.201709162
Viotti C, Krüger F, Krebs M, Neubert C, Fink F, Lupanga U, Scheuring D, Boutté Y, Frescattada-Rosa M, Wolfenstetter S, et al. (2013) The endoplasmic reticulum is the main membrane source for biogenesis of the lytic vacuole in Arabidopsis. Plant Cell 25: 3434–3449
Zhu L, Jorgensen J, Li M, Chuang Y-S, Emr SD (2017) ESCRTs function directly on the lysosome membrane to downregulate ubiquitinated lysosomal membrane proteins. eLife 6: e26403