Tools to unravel the mystery of why some tomatoes are hairier than others
Galdon-Armero et al. present an image resource that can be used to identify natural variation in leaf epidermal development in tomato. The Plant Cell (2020). https://doi.org/10.1105/tpc.20.00127
By Javier Galdon-Armero and Cathie Martin, John Innes Centre, Norwich, UK
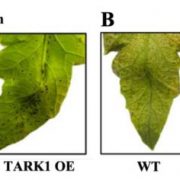
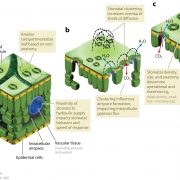
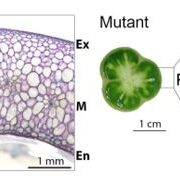
Background: Just like our skin, the plant epidermis provides an interface with the environment and protects against biotic and abiotic stresses. Hairs (trichomes) on the epidermis reduce heat load and water loss, while also providing defence chemicals that repel insect pests. Solanum pennellii is a wild tomato species endemic to arid habitats of the Andean regions in South America. In contrast, the domesticated tomato Solanum lycopersicum thrives in temperate, well-irrigated conditions. Introgression lines (ILs) between these two species have defined a wealth of natural variation for various traits. There are seven different types of trichomes described for the genus Solanum, both glandular and aglandular, but many are not easily visible by eye or with light microscopy. A high resolution analysis of epidermal cell development in the ILs is lacking.
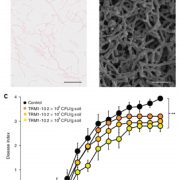
Question: We performed a comprehensive analysis of the leaf epidermis of two generations of the S. lycopersicum x S. pennellii ILs by a combination of Scanning Electron Microscopy (SEM) techniques. The intention was to provide a large data resource for further research into cellular development in tomato and related Solanaceae.
Findings: The image resource led to the identification of unexplored genomic regions associated with the determination of stomatal and trichome density in leaves, as well as trichome morphogenesis. To demonstrate how this unique resource can be used, we identified a new function of a transcription factor (previously described to control conical cell formation) in determining trichome density in leaves. The identification of natural variation governing epidermal development should allow breeding for improved epidermal traits, such as improved trichome density conferring improved water use efficiency, using the extensive resources available in tomato.
Next steps: The synteny between the tomato genome and important closely related crops like potato and eggplant as well as with more distant relatives in the Solanaceae family like pepper and tobacco means that this mapped resource of natural variation in trichome development could be indicative of genomic regions conferring effects on trichome formation in these other species, which will be of importance in identifying traits impacting drought tolerance and insect resistance, amongst others.
Galdon-Armero, et al. (2020). A Scanning Electron Micrograph-based Resource for Identification of Loci Involved in Epidermal Development in Tomato: Elucidation of a New Function for the Mixta-like Transcription Factor in Leaves. Plant Cell DOI: https://doi.org/10.1105/tpc.20.00127