The Evening Complex of the Circadian Clock and Chromatin
The circadian clock generates endogenous biological rhythms with periods of ~24 h. Circadian transcription allows the molecular anticipation of the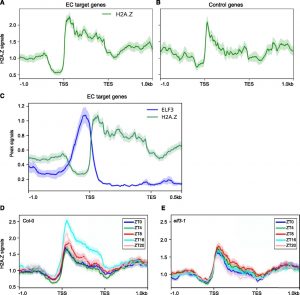 environmental cycles, which improves plant fitness and adaptation. Consistent with its adaptive function, the circadian clock is subject to multiple layers of regulation, which contribute toward accurate oscillations. A large fraction of the Arabidopsis (Arabidopsis thaliana) transcriptome is circadian-regulated. The Arabidopsis central oscillator consists of a series of sequential regulatory loops composed of genes expressed at different times during the diurnal cycle. Certain morning-expressed genes, for example, repress the expression of evening-expressed genes and vice versa. Among the repressors of morning gene expression are components of the Evening Complex (EC), which represses target gene expression at the end of the day and integrates temperature information to coordinate environmental and endogenous signals. Tong et al. (10.1104/pp.19.00881) now show that the EC induces repressive chromatin restructuring to regulate the evening transcriptome. The EC component EARLY FLOWERING3 (ELF3) directly interacts with a protein from the SWI2/SNF2-RELATED (SWR1) complex that is responsible for deposition of an important histone variant H2A.Z that has multiple effects on transcription. SWR1 components display circadian oscillation in gene expression with a peak at dusk. In turn, SWR1 is required for the circadian clockwork, since defects in SWR1 activity alter morning-expressed genes. The authors also show that the EC-SWR1 complex binds to the loci of the core clock genes PSEUDO-RESPONSE REGULATOR7 (PRR7) and PRR9. These results indicate that diurnal H2A.Z deposition contributes to circadian gene expression in Arabidopsis.
environmental cycles, which improves plant fitness and adaptation. Consistent with its adaptive function, the circadian clock is subject to multiple layers of regulation, which contribute toward accurate oscillations. A large fraction of the Arabidopsis (Arabidopsis thaliana) transcriptome is circadian-regulated. The Arabidopsis central oscillator consists of a series of sequential regulatory loops composed of genes expressed at different times during the diurnal cycle. Certain morning-expressed genes, for example, repress the expression of evening-expressed genes and vice versa. Among the repressors of morning gene expression are components of the Evening Complex (EC), which represses target gene expression at the end of the day and integrates temperature information to coordinate environmental and endogenous signals. Tong et al. (10.1104/pp.19.00881) now show that the EC induces repressive chromatin restructuring to regulate the evening transcriptome. The EC component EARLY FLOWERING3 (ELF3) directly interacts with a protein from the SWI2/SNF2-RELATED (SWR1) complex that is responsible for deposition of an important histone variant H2A.Z that has multiple effects on transcription. SWR1 components display circadian oscillation in gene expression with a peak at dusk. In turn, SWR1 is required for the circadian clockwork, since defects in SWR1 activity alter morning-expressed genes. The authors also show that the EC-SWR1 complex binds to the loci of the core clock genes PSEUDO-RESPONSE REGULATOR7 (PRR7) and PRR9. These results indicate that diurnal H2A.Z deposition contributes to circadian gene expression in Arabidopsis.