The Cold Never Bothered Me Anyway: DELLA-Interacting GROWTH REGULATING FACTORS Mediate Plant Growth in Cold Stress
The gibberellin (GA) hormone signaling pathway is a key regulator of major plant developmental processes such as seed germination, cell elongation, and flowering time. The influence of GA is mediated by DELLA repressor proteins which, in the absence of GA, inhibit the activity of transcription factors. In addition to general developmental processes, GA signaling is also vital for growth response to abiotic stressors such as cold temperature (Van De Velde et al., 2017). However, the multitude of molecular cold response pathways makes it difficult to determine mechanistic relationships using genetic approaches (Ding et al., 2019). Therefore, the mechanisms underlying DELLA-mediated cold stress regulation are currently unclear. In this issue of The Plant Cell, Lantzouni et al. (2020) combine RNA-seq with a yeast two-hybrid (Y2H) approach to identify GROWTH REGULATING FACTORS (GRFs) as DELLA targets with roles in GA-dependent plant growth control during cold stress.
 Since cold exposure broadly affects many physiological processes, it has been challenging to phenotypically disentangle the role of GA in plant growth responses. However, DELLAs accumulate early in response to cold temperatures, prior to phenotypic growth changes. Thus, using a molecular approach, Lantzouni and colleagues tested global transcriptional changes resulting from cold stress and GA treatment of Arabidopsis seedlings. The data support a feedback system whereby cold stress controls DELLA abundance via the regulation of GA biosynthesis and signaling genes. Further, combining the cold and GA treatment RNA-seq datasets provides insight into the contribution of DELLAs to cold stress-induced gene expression changes. Less than a quarter of genes that were GA-regulated after cold stress were also GA-regulated at ambient temperature, suggesting differential transcription factor regulation by DELLAs at the two temperatures. Additionally, about 75% of the genes regulated by GA in cold treated samples were also differentially expressed in response to cold stress alone, pointing to an active role for DELLAs in the regulation of the transcriptional cold response via integration with the cold stress pathway.
Since cold exposure broadly affects many physiological processes, it has been challenging to phenotypically disentangle the role of GA in plant growth responses. However, DELLAs accumulate early in response to cold temperatures, prior to phenotypic growth changes. Thus, using a molecular approach, Lantzouni and colleagues tested global transcriptional changes resulting from cold stress and GA treatment of Arabidopsis seedlings. The data support a feedback system whereby cold stress controls DELLA abundance via the regulation of GA biosynthesis and signaling genes. Further, combining the cold and GA treatment RNA-seq datasets provides insight into the contribution of DELLAs to cold stress-induced gene expression changes. Less than a quarter of genes that were GA-regulated after cold stress were also GA-regulated at ambient temperature, suggesting differential transcription factor regulation by DELLAs at the two temperatures. Additionally, about 75% of the genes regulated by GA in cold treated samples were also differentially expressed in response to cold stress alone, pointing to an active role for DELLAs in the regulation of the transcriptional cold response via integration with the cold stress pathway.
In a complementary approach, a Y2H assay identified over 250 structurally diverse DELLA-interacting transcription factors. To uncover novel DELLA interactors that function in GA-regulated cold responses, the authors compared the Y2H and RNA-seq datasets to search for DELLA-interacting transcription factors that are also GA-regulated at the transcriptional level during cold stress. Notably, six of the nine Arabidopsis GRF transcription factor family members interacted with DELLAs and the transcription of three was GA-regulated in response to cold stress. GRFs are known to regulate plant growth, particularly in leaves and roots, and recent work biochemically verified in vivo interactions between GRFs and DELLAs in rice (Li et al., 2018). Excitingly, Arabidopsis plants with altered GRF levels are differentially sensitive to the manipulation of GA and, as a result, DELLA levels, with respect to root and leaf growth. Together, the data indicate that GA and subsequently DELLAs regulate organ growth through GRFs.
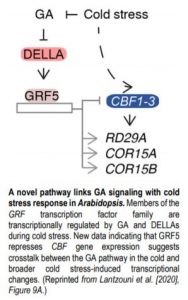
To identify genes that are downstream of DELLA-mediated GRF5 regulation, an additional RNA-seq experiment profiled transcriptomic changes in response to both GA and cold stress in seedlings overexpressing GRF5. A result of particular interest is the 35S:GRF5-mediated attenuation of cold stress-dependent induction of all three C-Repeat BINDING FACTORs (CBFs), which are known regulators of cold responses. Further, the expression levels of cold-regulated genes such as RD29A, COR15A, and COR15B, which are known CBF targets, were upregulated. Together, the reported data indicate that GRFs repress CBF transcription and point to possible crosstalk between the GA pathway in the cold and general cold stress-induced gene expression changes (see Figure).
In summary, this work identifies GRFs as DELLA-mediated regulators of cold temperature-induced gene expression changes and uncovers a novel pathway linking GA signaling with cold stress response. The reported transcriptomic and interaction datasets are valuable resources for further molecular investigation.
Rachel Shahan
Department of Biology
Duke University
ORCID: 0000‐0001‐7702‐9432
REFERENCES
Ding Y, Shi Y and Yang S. (2019). Advances and challenges in uncovering cold tolerance regulatory mechanisms in plants. New Phytol. 222: 1690-1704.
Lantzouni O, Alkofer A, Falter-Braun P and Schwechheimer C. (2020). GROWTH-REGULATING FACTORS interact with DELLAs and regulate growth in cold stress. Plant Cell. Published February 2020. https://doi.org/10.1105/tpc.19.00784
Li S, Tian Y, Wu K, Ye Y, Yu J, Zhang J, Liu Q, Hu M, Li H, Tong Y, Harberd NP and Fu X. (2018). Modulating plant growth-metabolism coordination for sustainable agriculture. Nature 560: 595-600.
Van De Velde K, Ruelens P, Geuten K, Rohde A and Van der Straeten D. (2017). Exploiting DELLA Signaling in Cereals. Trends Plant Sci. 22: 880-893.