
New kid on the plant block: Single-cell proteomics
Plant Science Research WeeklyWhile single-cell omics technologies, particularly transcriptomics, are already becoming widely adopted in plant science, quantifying proteins at single cell resolution is less established. Fortunately, important technological strides have been made that improve sample preparation, separation techniques,…
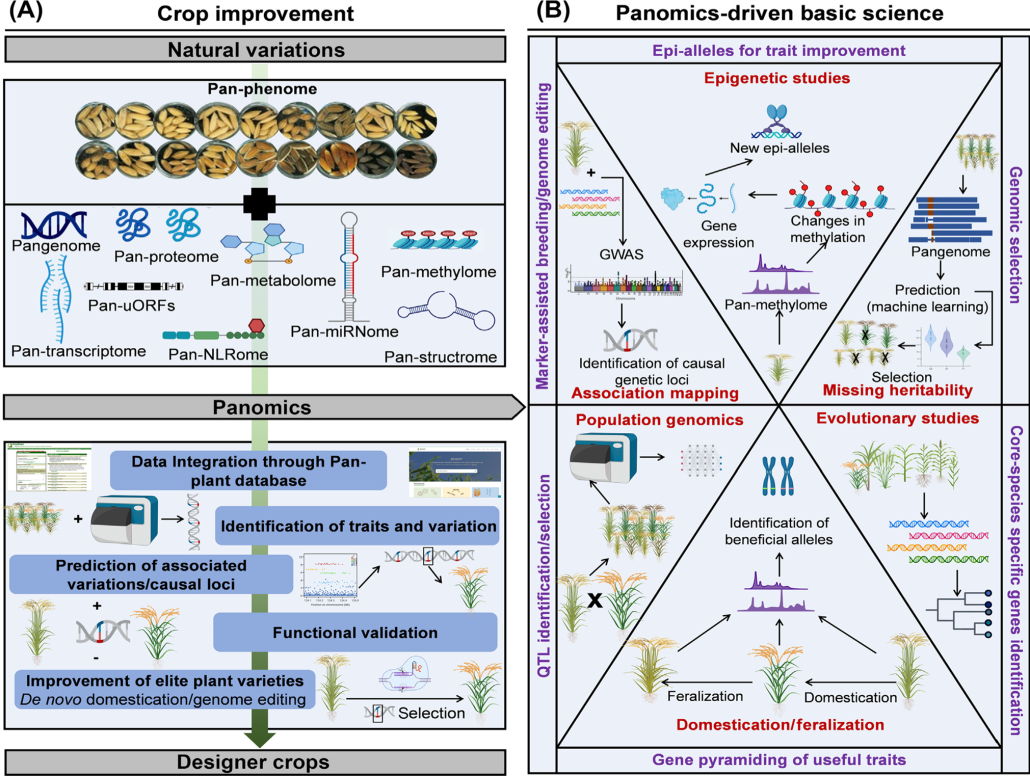
Review: The era of panomics-driven gene discovery in plants
Plant Science Research WeeklyPanomics, an approach integrating multiple ‘omics’ datasets such as genomics, transcriptomics, metabolomics, and phenomics, has seen rapid advancement in recent years due to technological improvements, particularly in genomics. This review focuses on the recent developments in panomics-driven gene…
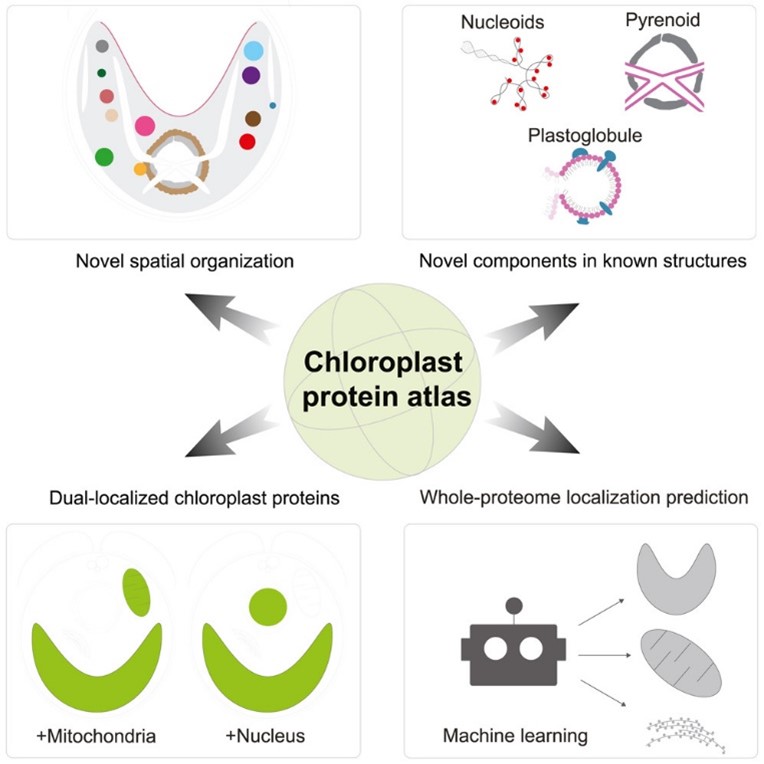
A chloroplast protein atlas reveals punctate structures and spatial organization of biosynthetic pathways
Plant Science Research WeeklyChloroplasts are the location of key processes including photosynthesis, starch synthesis and lipid synthesis. However, many chloroplast proteins have unknown functions, a problem that can in part by addressed through high-resolution localization data. Here, 1034 putative chloroplast- localized proteins…
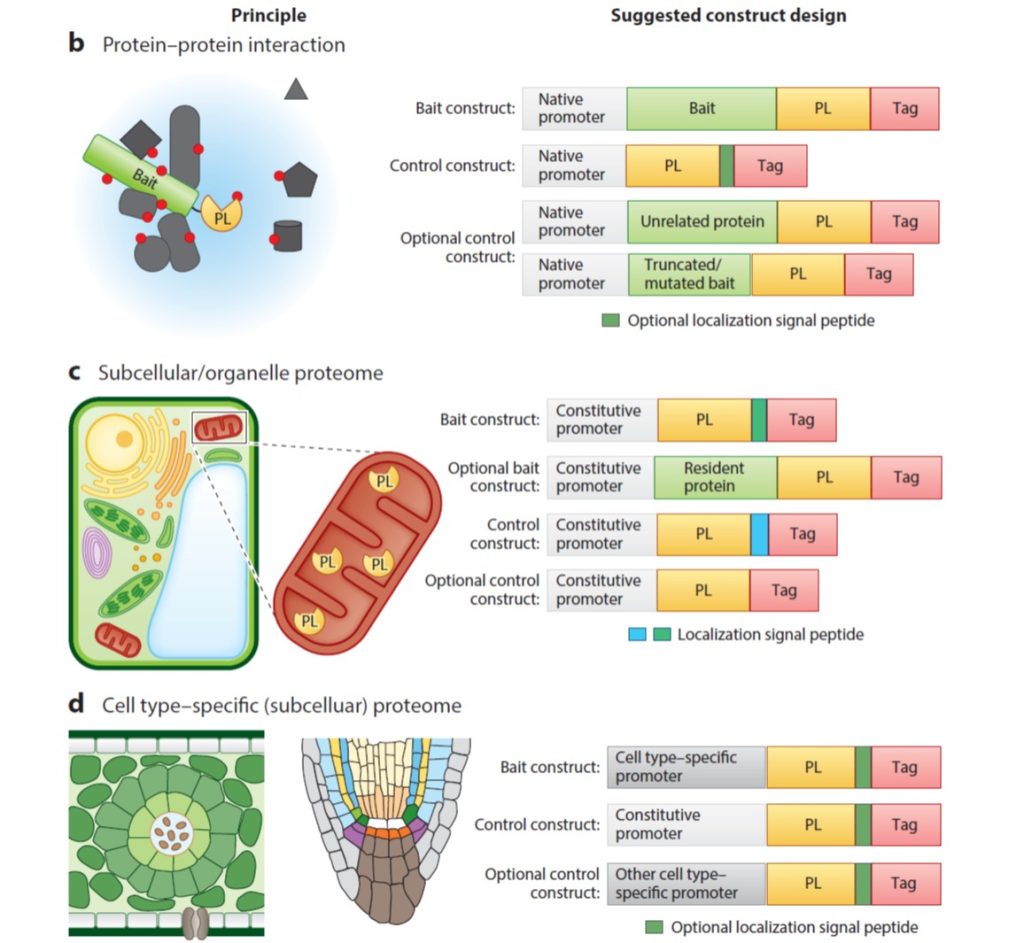
Review: Proximity labeling in plants
Plant Science Research WeeklyGenetic studies can suggest that two proteins function in the same pathway, but how can we figure out if they share the same space? In this review, Xu et al. provide an overview of proximity labeling, a method to identify proteins that co-localize in space. Proximity labeling uses a biotin ligase which…
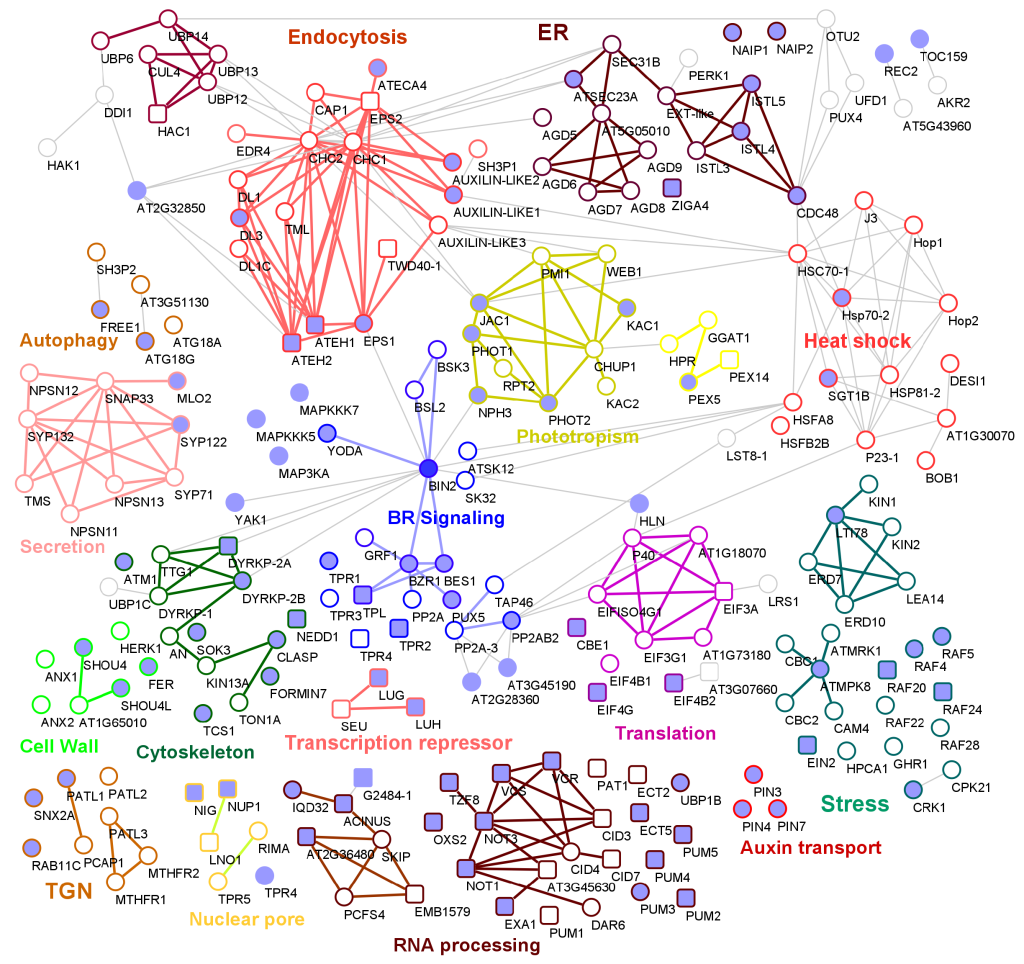
BIN2 signaling network constructed using proximity labeling and TurboID
Plant Science Research WeeklyThe brassinosteroid (BR) signaling pathway is essential for plant growth and development. BRASSINOSTEROID-INSENSITIVE2 (BIN2) is a GSK3-like kinase, serving as a repressor of the BR signaling pathway. In the absence of BR, BIN2 phosphorylates BRASSINAZOLE-RESISTANT1 (BZR1) family transcription factors…
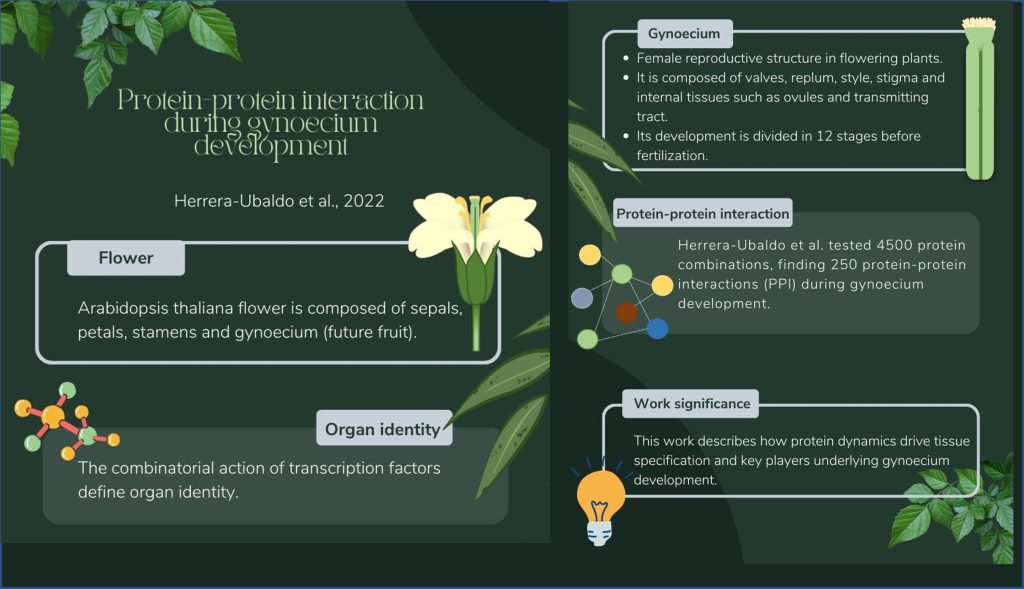
Protein-protein interaction landscape of transcription factors during gynoecium development in Arabidopsis (Mol. Plant)
Plant Science Research WeeklyThe gynoecium is the female reproductive structure of flowering plants. This complex organ, composed of different tissues, is an excellent model for studying plant organ development. Although the gynoecium has been extensively studied from a genetic point of view and a few studies have been carried out…
A Contextualized Catalog: Proteomic analyses of plant CCVs
The Plant Cell: In a NutshellDahhan et al. explore the clathrin-coated vesicle proteome in Arabidopsis
By Dana A. Dahhan and Sebastian Y. Bednarek, Dept. of Biochemistry, UW-Madison
Background: Trafficking of vacuolar and plasma membrane proteins and other macromolecules is essential for plant development, nutrient uptake,…
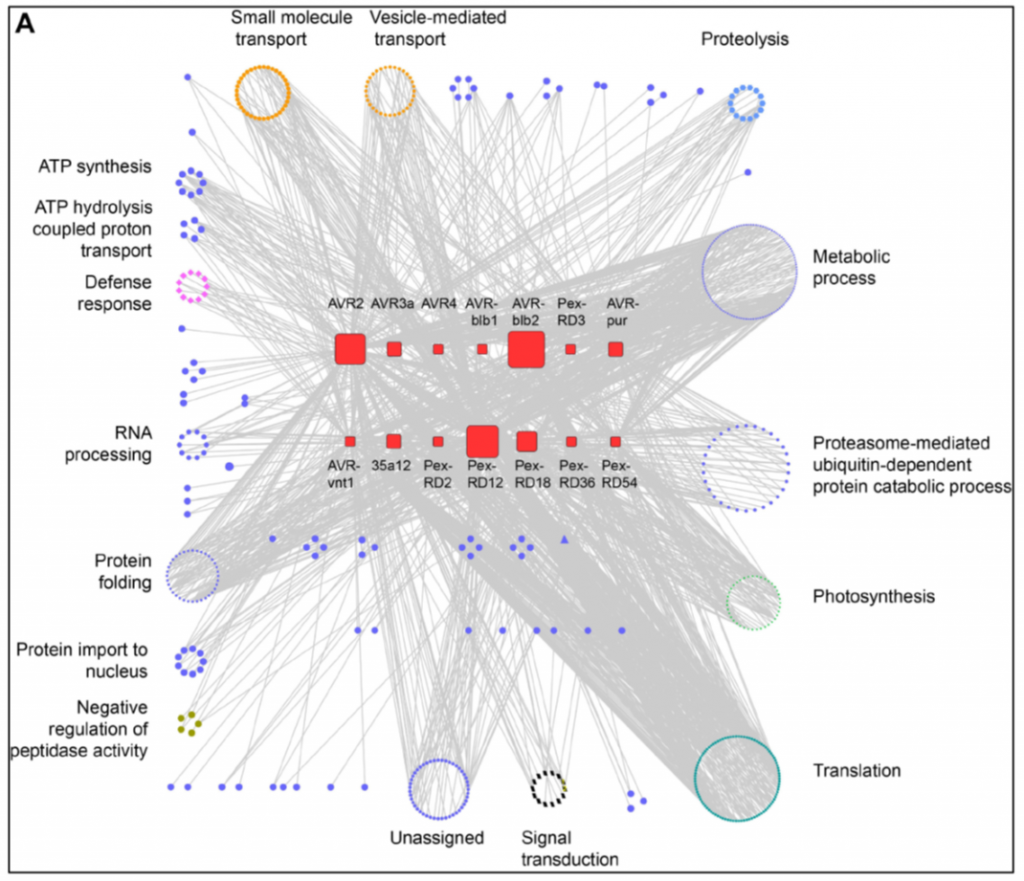
Host-interactor screens of Phytophthora infestans RXLR proteins reveal vesicle trafficking as a major effector-targeted process (Plant Cell)
Plant Science Research WeeklyIt’s like the plot of every spy movie you’ve ever seen. If you could infiltrate your opponent’s headquarters what would you target to disable your enemy? This is the question addressed in new work by Petre et al. More than ten years ago, a family of small proteins was identified, the RXLR proteins…
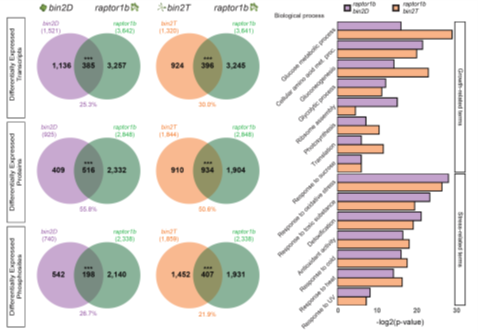
Multi-omics analysis reveals interplay between BR and TORC signaling (bioRxiv)
Plant Science Research WeeklyPlants have evolved well-coordinated crosstalk between different signaling pathways to respond and adapt to various environmental stresses. Brassinosteriods (BRs) and Target of Rapamycin Complex (TORC) have multiple roles, through transcription, translation and autophagy, to control the balance between…

