Selection Drives Gene Escape from Centromeres
Liao et al. performed comparative phylogenomic analysis on rice centromeres. Plant Cell (2018). https://doi.org/10.1105/tpc.18.00163
By Yi Liao and Mingsheng Chen
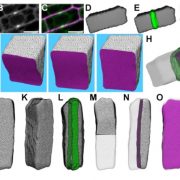
Background: Centromeres are necessary for faithful chromosome segregation in eukaryotic organisms. The genomic regions conferring centromere function/identity in most plants and animals are composed of large arrays of satellite repeats, surrounded by other highly repetitive DNA sequences. During evolution, centromere locations can change through chromosomal arrangements and some unclear epigenetic mechanisms such as centromere repositioning. Both centromeric chromatin and flanking pericentromeric heterochromatin are thought to be incompatible with gene transcription and thus represent an unfriendly environment for gene survival. However, centromeres may evolve from gene-containing regions and active genes have indeed been found within centromeric regions.
Question: How and to what extent have centromere locations changed and what is the evolutionary fate of genes within centromeric and pericentromeric regions? We study these questions in rice (Oryza spp) in which abundant high-quality genome sequences are available, facilitating comparative phylogenomic analysis.
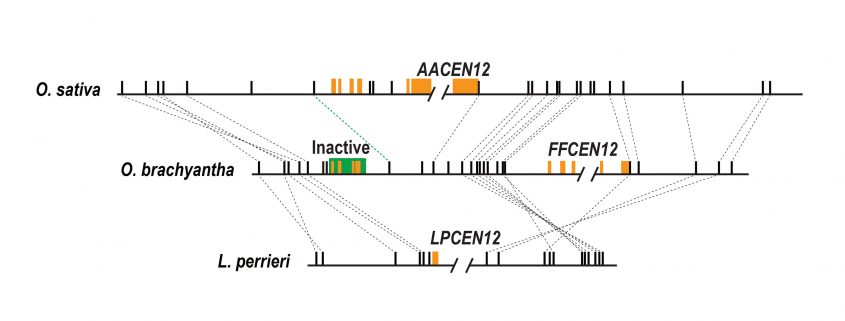
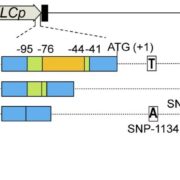
Findings: We found that the physical positions of the centromeres in Oryza sativa and Oryza brachyantha, species which diverged approximately 15 million years ago, are generally conserved. Minor alterations in their centromere locations are due to local chromosomal arrangements (e.g., inversions) and/or centromere repositioning that can be triggered by duplicated transposition. We observed an excess of syntenic gene loss in the centromeric and pericentromeric regions of the rice genome. Most of the lost genes were found to have moved to other genomic regions through segmental duplications, suggesting an evolutionary trend of centromere-linked genes escaping from centromeric regions. The excess gene loss is consistent with selective loss of the parental gene copy in dynamic centromeric region after gene duplications and supports the antagonistic relationship between the centromere environment and gene survival. We also observed abundant new genes at pericentromeric regions.
Next steps: Our study showed that selective forces conferred by centromere chromatin dynamics may drive gene relocation out of centromeric regions. It will be interesting to investigate how new genes evolve and fix at pericentromeric regions.
Yi Liao, Xuemei Zhang, Bo Li, Tieyan Liu, Jinfeng Chen, Zetao Bai, Meijiao Wang, Jinfeng Shi, Jason G. Walling, Rod A. Wing, Jiming Jiang, Mingsheng Chen. (2018). Comparison of Oryza sativa and Oryza brachyantha Genomes Reveals Selection-Driven Gene Escape from the Centromeric Regions. Plant Cell 30: 1729-1744; https://doi.org/10.1105/tpc.18.00163