Rapid changes: ABA-independent SnRK2s target mRNA decay
Magda Julkowska
Magdalena.Julkowska@kaust.edu.sa
In response to stress, secondary messengers and rapid and reversible protein phosphorylation contribute to signaling cascades that generate unique signatures indicating stress type and severity. Activation of stress-induced signaling cascades leads to changes in the transcriptome and ultimately to acclimatization to stress conditions. Changes in the transcriptome result not only from the altered activity of transcription factors, but also from the regulation of cytoplasmic mRNA dynamics.
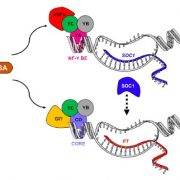
The fate of cytoplasmic mRNA is coordinated by heterogenous mRNA-ribonucleoprotein complexes (mRNP), including polyribosomes, processing bodies, and stress granules(Chantarachot and Bailey-Serres 2018). The core components of processing bodies include VCS (VARICOSE), DCP1 (DECAPPING PROTEIN 1) and DCP2. DCP2 removes the 5′ m7G cap from the mRNA, thereby inhibiting translation initiation. The formation of the complex consisting of DCP2, DCP1 and VCS is necessary for decapping activity (Xu et al. 2006). While mRNA without a m7G cap is more likely to undergo degradation by being targeted with EXORIBONUCLEASE4 (XRN4), some processing-body targeted mRNA has been reported to be stabilized, rather than degraded (Scarpin et al. 2017), illustrating the context-dependent fate of mRNA targeted to mRNPs. Mutant alleles of processing body core components show strong defects in post-embryonic development (Xu et al. 2006), making it difficult to study how environmental signals modify the transcriptome through processing bodies.
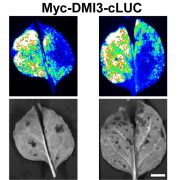
Previously, VCS has been identified as a phosphorylation target of abscisic acid (ABA)-independent protein kinases belonging to the SnRK2 (Sucrose non-fermenting-1 related Protein Kinase2s) family (Soma et al. 2017). In this issue of Plant Physiology, a study by Kawa (et al. 2019) confirms that under salt stress, ABA-independent SnRK2s phosphorylate core components of the mRNA decapping complex. The group mapped the phosphorylation sites and described how the changes in transcriptome contribute to changes in root architecture. Upon salt stress, ABA-independent SnRK2.4 and SnRK2.10 are rapidly activated (maximum activity between 30 sec. and 5 minutes), and relocate into punctate structures (McLoughlin et al. 2012). By examining the co-purified proteins using tandem-affinity purification for SnRK2.4 and SnRK2.10, Kawa (et al. 2019) identified that SnRK2.4/2.10 bind and phosphorylate VCS, while DCP2 was part of the pulled-down complex, possibly by its binding to VCS.
By studying the transcriptome of the SnRK2 mutant lines, Kawa (et al., 2019) identified that salt-induced transcriptional changes in genes associated with lateral root development, constituting the substrates of the VCS, are absent from the SnRK2 mutant lines. One of these genes encodes a cytochrome P450 (CYP79B2), which participates in the formation of auxin via indole-3-acetaldoxime (Mashiguchi et al. 2011). The salt stress results in induced transcription of CYP79B2, previously associated with the maintenance of lateral root development under salt stress (Julkowska et al. 2017). However, this increase in expression of CYP79B2 is reduced in SnRK2 mutants (Kawa et al., 2019). The reduction in lateral root development upon salt stress was previously observed in mutants of SnRK2.4 (McLoughlin et al. 2012), similarly to CYP79B2 mutants (Julkowska et al. 2017). While it remains to be determined how phosphorylation of the core components of the mRNA decay pathway affects the fate of mRNA, the study by Kawa (et al., 2019) provides an exciting perspective on how protein kinase activity can affect the transcriptional landscape, leading to a rapid adjustment in plant development.
References:
Chantarachot, Thanin, and Julia Bailey-Serres. 2018. “Polysomes, Stress Granules, and Processing Bodies: A Dynamic Triumvirate Controlling Cytoplasmic mRNA Fate and Function.” Plant Physiology 176 (1): 254–69.
Julkowska, Magdalena M., Iko T. Koevoets, Selena Mol, Huub Hoefsloot, Richard Feron, Mark A. Tester, Joost J. B. Keurentjes, et al. 2017. “Genetic Components of Root Architecture Remodeling in Response to Salt Stress.” The Plant Cell 29 (12): 3198–3213.
Kawa, Dorota, A. Jessica Meyer, Henk L. Dekker, Ahmed Abd-El-Haliem, Kris Gevaert, Eveline Van De Slijke, Justyna Maszkowska, et al. 2019. “SnRK2 Protein Kinases and mRNA Decapping Machinery Control Root Development and Response to Salt.” Plant Physiology, September. https://doi.org/10.1104/pp.19.00818.
Mashiguchi, Kiyoshi, Keita Tanaka, Tatsuya Sakai, Satoko Sugawara, Hiroshi Kawaide, Masahiro Natsume, Atsushi Hanada, et al. 2011. “The Main Auxin Biosynthesis Pathway in Arabidopsis.” Proceedings of the National Academy of Sciences of the United States of America 108 (45): 18512–17.
McLoughlin, Fionn, Carlos S. Galvan-Ampudia, Magdalena M. Julkowska, Lotte Caarls, Dieuwertje van der Does, Christiane Laurière, Teun Munnik, Michel A. Haring, and Christa Testerink. 2012. “The Snf1-Related Protein Kinases SnRK2. 4 and SnRK2. 10 Are Involved in Maintenance of Root System Architecture during Salt Stress.” The Plant Journal: For Cell and Molecular Biology 72 (3): 436–49.
Scarpin, María R., Lorena Sigaut, Silvio G. Temprana, Graciela L. Boccaccio, Lía I. Pietrasanta, and Jorge P. Muschietti. 2017. “Two Arabidopsis Late Pollen Transcripts Are Detected in Cytoplasmic Granules.” Plant Direct 1 (4): e00012.
Soma, Fumiyuki, Junro Mogami, Takuya Yoshida, Midori Abekura, Fuminori Takahashi, Satoshi Kidokoro, Junya Mizoi, Kazuo Shinozaki, and Kazuko Yamaguchi-Shinozaki. 2017. “ABA-Unresponsive SnRK2 Protein Kinases Regulate mRNA Decay under Osmotic Stress in Plants.” Nature Plants 3 (January): 16204.
Xu, Jun, Jun-Yi Yang, Qi-Wen Niu, and Nam-Hai Chua. 2006. “Arabidopsis DCP2, DCP1, and VARICOSE Form a Decapping Complex Required for Postembryonic Development.” The Plant Cell 18 (12): 3386–98.