Moonlighting Enzymes: How Often Are We Missing Secondary Functions?
We think of enzymes as highly specific catalysts that carry out one reaction and show nearly absolute substrate specificity. However, absolute specificity is the exception, not the rule, as most enzymes accept several structurally similar substrates. Moreover, many enzymes catalyze alternative reactions and, in some cases, even perform functions that are best described as nonenzymatic (Moore, 2004; Huberts and van der Klei, 2010). These alternative functions are sometimes referred to as moonlighting activity. A recent example of such an enzyme with a nonenzymatic function was published in this issue of Plant Physiology (Schmid et al., 2018).
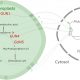
 The authors showed that the nuclear-encoded plastid UMP kinase from Arabidopsis (Arabidopsis thaliana), PUMPKIN, catalyzes the phosphorylation of UMP to UDP with the cosubstrate ATP. While the kinetic parameters are similar to the characterized homolog from E. coli, the loss of enzymatic function alone in transfer DNA insertion and RNAi lines could not explain the phenotype of severe growth retardation, photobleaching, and an increase in generation time (∼21 weeks versus ∼ 8 weeks for the wild type). Through quantitation of major plastid proteins and analysis of de novo protein synthesis rates, the authors showed that the loss of PUMPKIN affects protein translation in plastids. They followed this up with experiments that show that PUMPKIN is a compound of several large protein complexes in the plastid, of which some are sensitive to RNase treatment.
The authors showed that the nuclear-encoded plastid UMP kinase from Arabidopsis (Arabidopsis thaliana), PUMPKIN, catalyzes the phosphorylation of UMP to UDP with the cosubstrate ATP. While the kinetic parameters are similar to the characterized homolog from E. coli, the loss of enzymatic function alone in transfer DNA insertion and RNAi lines could not explain the phenotype of severe growth retardation, photobleaching, and an increase in generation time (∼21 weeks versus ∼ 8 weeks for the wild type). Through quantitation of major plastid proteins and analysis of de novo protein synthesis rates, the authors showed that the loss of PUMPKIN affects protein translation in plastids. They followed this up with experiments that show that PUMPKIN is a compound of several large protein complexes in the plastid, of which some are sensitive to RNase treatment.
RNA-immunoprecipitation combined with deep sequencing and additional experiments convincingly showed that PUMPKIN associates with group II introns of five genes of the chloroplast genome. PUMPKIN stabilizes its target RNAs but does not seem to be involved in splicing of these introns. Interestingly, a small number of group II introns are catalytically active ribozymes that self-excise from their mRNA in vitro; however, in vivo proteins are usually required for splicing of group II introns (Bonen and Vogel, 2001). The mechanism by which PUMPKIN stabilizes its RNA targets is unknown, and no known RNA-binding motifs were found in PUMPKIN. Therefore, the identity of the RNA sequence and/or 3D structure that is bound remains a target of future research. The authors speculate that multimerization of PUMPKIN could explain RNA binding, given that its enzymatic function already involves binding of one RNA monophosphate nucleotide. Thus, individual subunits of a multimeric enzyme could bind an RNA sequence with high enough affinity.
In light of this research, we have to ask ourselves: how often are we overlooking moonlighting activities? And does moonlighting activity explain why in vitro biochemical characterizations cannot always be linked to mutant phenotypes?
REFERENCES
Bonen L, Vogel J (2001) The ins and outs of group II introns. Trends Genet 17: 322–331
Huberts DH, van der Klei IJ (2010) Moonlighting proteins: an intriguing mode of multitasking. Biochim Biophys Acta 1803: 520–525
Moore Bd (2004) Bifunctional and moonlighting enzymes: lighting the way to regulatory control. Trends Plant Sci 9: 221–228
Schmid L-M, Ohler L, Möhlmann T, Brachmann A, Muiño J, Leister D, Meurer J, Manavski N (2018) PUMPKIN, the sole plastid UMP Kinase associates with Group II Introns and impacts their metabolism. Plant Physiology 179: 248–264 10.1104/pp.18.00687