Get it Sorted: A Classic Endocytic Sorting Mechanism in Mammals is Conserved in Plants
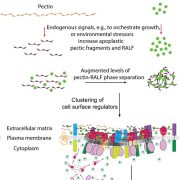
The plasma membrane (PM) is an exquisitely dynamic structure. It has to be, since it serves as the interface between the cell and the ever-changing environment. PM-associated signaling proteins shuttle quickly between the PM and the cell’s interior to help the organism respond to whatever comes its way. Cells employ pathways such as clathrin-mediated endocytosis (CME) to selectively engulf a target protein in the PM and move it into the cell as needed to fine-tune signaling (reviewed in Reynolds et al., 2018). In the mammalian CME pathway, PM proteins marked by specific tyrosine or di-leucine motifs in their cytoplasmic tails are recognized by the heterotetrameric adaptor protein complex-2 (AP-2). AP-2 recruits lattice-like clathrin molecules, which coat the budding PM and form a cage around the cargo-packed vesicle that carries the protein to its destination. The most common motif, YXXΦ (where Y is tyrosine, X is any amino acid, and Φ is a bulky hydrophobic residue), is recognized by the medium AP-2 subunit AP2M.
 Although the CME pathway plays crucial roles in modulating plant physiology, very little is known about how PM proteins in plants are sorted by AP-2 for CME and whether YXXΦ motifs are required for protein recognition. To probe this issue, Derui Liu and colleagues (Liu et al., 2020) studied CME of the brassinosteroid (BR) receptor BR INSENSITIVE1 (BRI1) in Arabidopsis thaliana, a process known to depend on clathrin and AP-2. BRs play multiple roles throughout the plant lifecycle (reviewed in Nolan et al., 2020). The PM pool of BRI1 controls BR signaling: impaired endocytosis of this receptor kinase leads to constitutive BR responses. To look for interactions between BRI1 and AP-2, the authors examined the dynamics of GFP-tagged BRI1 and RFP-tagged large AP-2 subunit AP2A1 in the PM of root epidermal cells of the null mutant bri1-116. A portion of BRI1 and AP2A1 displayed similar dynamics and disappeared from the PM together. The direct interaction between BRI1 and AP2M was confirmed in vitro.
Although the CME pathway plays crucial roles in modulating plant physiology, very little is known about how PM proteins in plants are sorted by AP-2 for CME and whether YXXΦ motifs are required for protein recognition. To probe this issue, Derui Liu and colleagues (Liu et al., 2020) studied CME of the brassinosteroid (BR) receptor BR INSENSITIVE1 (BRI1) in Arabidopsis thaliana, a process known to depend on clathrin and AP-2. BRs play multiple roles throughout the plant lifecycle (reviewed in Nolan et al., 2020). The PM pool of BRI1 controls BR signaling: impaired endocytosis of this receptor kinase leads to constitutive BR responses. To look for interactions between BRI1 and AP-2, the authors examined the dynamics of GFP-tagged BRI1 and RFP-tagged large AP-2 subunit AP2A1 in the PM of root epidermal cells of the null mutant bri1-116. A portion of BRI1 and AP2A1 displayed similar dynamics and disappeared from the PM together. The direct interaction between BRI1 and AP2M was confirmed in vitro.
The cytoplasmic domain of BRI1 contains five putative surface-exposed YXXΦ endocytic motifs. After narrowing the list to three candidates, the authors examined their functions in vivo by generating full-length BRI1 carrying mutations in each motif, C-terminally tagged the mutant proteins with mCitrine, and expressed them in the Arabidopsis bri1 null allele. All three mutant proteins complemented bri1. However, only plants with a tyrosine-to-phenylalanine substitution in BRI1 (BRI1Y898F) were hypersensitive to BRs, as revealed in a hypocotyl growth assay. Unlike the other mutations, the BRI1Y898F mutation did not impair the kinase activity of BRI1.
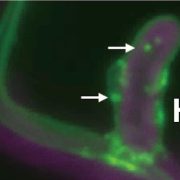
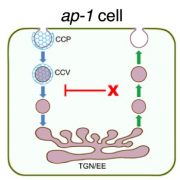
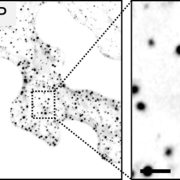
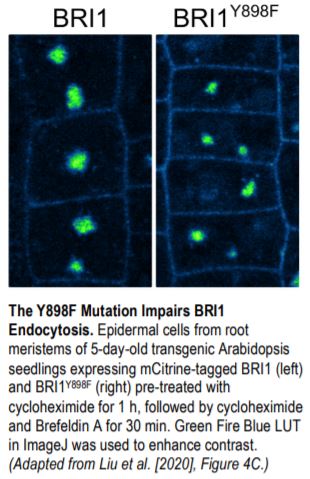
BR treatment leads to dephosphorylation of the transcription factor BES1, a commonly used indicator of BR signaling activation. Upon exogenous BR treatment, dephosphorylated BES1 accumulated at higher levels in BRI1Y898F vs. BRI1 plants, confirming the hypersensitivity of these plants to BRs. To determine whether this hypersensitivity was due to increased levels of BRI1 at the PM, mCitrine-tagged BRI1Y898F vs. BRI1 signals at the PM of root meristem cells were examined in transgenic bri1 seedlings following treatment with cycloheximide to inhibit de novo protein synthesis. BRI1Y898F generated significantly stronger fluorescent signals at the PM than BRI1. Brefeldin A (BFA) treatment to inhibit exocytosis led to fewer and smaller BFA bodies in plants harboring BRI1Y898F vs. BRI1 (see figure), indicating that endocytosis of the mutant protein was impaired. Finally, BRI1Y898F showed reduced binding to AP2M in an in vitro pull-down assay, confirming that AP2M recognizes this motif to facilitate CME.
As shown in the figure, disrupting the YXXΦ motif in BRI1 failed to fully block its internalization, raising the question of whether an additional sorting mechanism is at play. Stay tuned.
Jennifer Lockhart
Science Editor
ORCID: 0000-0002-1394-8947
REFERENCES
Nolan, T., Vukašinović, N., Liu, D., Russinova, E., and Yin, Y. (2020). Brassinosteroids: Multi-dimensional regulators of plant growth, development, and stress responses. Plant Cell 32: 295-318.
Reynolds, G.D., Wang, C., Pan, J., and Bednarek, S.Y. (2018). Inroads into internalization: five years of endocytic exploration. Plant Physiol. 176: 208-218.