Genomic diversification of LATE EMBRYOGENESIS ABUNDANT (LEA) protein gene families (GBE)
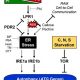
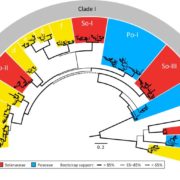
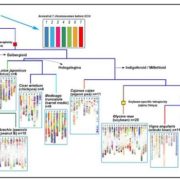
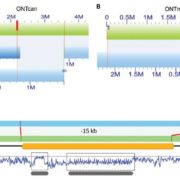
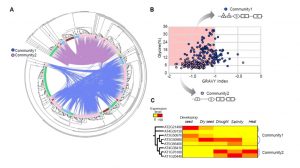 LEA genes were first identified as being highly abundant during seed desiccation (hence their name), but later were also shown to accumulate in other tissues in response to drought stress, and to confer desiccation tolerance in “resurrection plants”. These small proteins are characterized by having intrinsically disordered domains, shown to contribute to their properties of protecting macromolecular structures. LEA proteins are encoded by eight multi-gene families, raising questions about evolutionary origins and functional redundancies. Artur et al. explored LEA genes across 60 complete plant genomes, including desiccation tolerant angiosperms, nonvascular plants and green algae. Their findings suggest that two gene families predate terrestrial plants, but there has been extensive amplification in the terrestrial lineages, leading to diverse functions and expression patterns. Synteny analysis of LEA genes indicates that genes that are retained in chromosomal position across plant lineages are likely to have conserved function, and that synteny diversification is often correlated with functional diversification. (Summary by Mary Williams) Genome Biology and Evolution 10.1093/gbe/evy248.
LEA genes were first identified as being highly abundant during seed desiccation (hence their name), but later were also shown to accumulate in other tissues in response to drought stress, and to confer desiccation tolerance in “resurrection plants”. These small proteins are characterized by having intrinsically disordered domains, shown to contribute to their properties of protecting macromolecular structures. LEA proteins are encoded by eight multi-gene families, raising questions about evolutionary origins and functional redundancies. Artur et al. explored LEA genes across 60 complete plant genomes, including desiccation tolerant angiosperms, nonvascular plants and green algae. Their findings suggest that two gene families predate terrestrial plants, but there has been extensive amplification in the terrestrial lineages, leading to diverse functions and expression patterns. Synteny analysis of LEA genes indicates that genes that are retained in chromosomal position across plant lineages are likely to have conserved function, and that synteny diversification is often correlated with functional diversification. (Summary by Mary Williams) Genome Biology and Evolution 10.1093/gbe/evy248.