Field-based species identification of closely-related plants using real-time nanopore sequencing
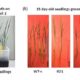
 DNA sequencing was slow before the development of high throughput sequencing. Portable DNA sequencing, which would make sequencing on-site a reality, was impossible until recently. Parker et al. report on the on-site use of MinION from Oxford Nanopore Technologies for DNA barcoding, which yields data within hours, showing “species identification could be conducted using genome scale data generated at the point of sample collection.” The paper describes the procedures that were followed to identify and distinguish samples from Arabidopsis thaliana and Arabidopsis lyrata, two closely related species. Their results demonstrate that “the entire process (from sample collection to genome scale sequencing) is now feasible for eukaryotic species within a few hours in field conditions.” Even with the caveats of lower sample purity and sequencing coverage, their data were useful for genome assembly and would also allow quick evolutionary analyses. As sequencing technology advances, it will be more and more reachable, even for non-experts, incredibly extending its uses—this is a nice time to do field work. Scientific Reports: 10.1038/s41598-017-08461-5 (Contributed by Gabriela Auge)
DNA sequencing was slow before the development of high throughput sequencing. Portable DNA sequencing, which would make sequencing on-site a reality, was impossible until recently. Parker et al. report on the on-site use of MinION from Oxford Nanopore Technologies for DNA barcoding, which yields data within hours, showing “species identification could be conducted using genome scale data generated at the point of sample collection.” The paper describes the procedures that were followed to identify and distinguish samples from Arabidopsis thaliana and Arabidopsis lyrata, two closely related species. Their results demonstrate that “the entire process (from sample collection to genome scale sequencing) is now feasible for eukaryotic species within a few hours in field conditions.” Even with the caveats of lower sample purity and sequencing coverage, their data were useful for genome assembly and would also allow quick evolutionary analyses. As sequencing technology advances, it will be more and more reachable, even for non-experts, incredibly extending its uses—this is a nice time to do field work. Scientific Reports: 10.1038/s41598-017-08461-5 (Contributed by Gabriela Auge)