A chromosome conformation capture ordered sequence of the barley genome
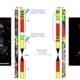
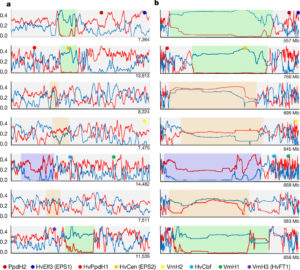 Cereal grasses are of course economically important, but they also have large repetitive genomes with large pericentromeric regions that have been difficult to map and sequence. Barley (Hordeum vulgare L.) is used for human and animal food and fermented to produce beer and whisky. A barley sequence assembly was published in 2012, but now Mascher et al. have used chromosome conformation capture mapping to produce a much more complete barley genome sequence, including sequences across the pericentromeric region. Their data reveal significant differences in gene density, recombination rates and retrotransposon distribution in the distal versus proximal regions of the chromosome. They also observe low levels of variation within elite germplasms, presumably due to intense selection. Nature 10.1038/nature22043
Cereal grasses are of course economically important, but they also have large repetitive genomes with large pericentromeric regions that have been difficult to map and sequence. Barley (Hordeum vulgare L.) is used for human and animal food and fermented to produce beer and whisky. A barley sequence assembly was published in 2012, but now Mascher et al. have used chromosome conformation capture mapping to produce a much more complete barley genome sequence, including sequences across the pericentromeric region. Their data reveal significant differences in gene density, recombination rates and retrotransposon distribution in the distal versus proximal regions of the chromosome. They also observe low levels of variation within elite germplasms, presumably due to intense selection. Nature 10.1038/nature22043




Leave a Reply
Want to join the discussion?Feel free to contribute!