1135 Arabidopsis genomes reveal global pattern of polymorphism
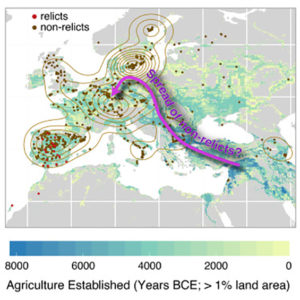 There are many accessions of Arabidopsis thaliana beyond the ecotypes predominantly used in research laboratories. In this article, The 1001 Genomes Consortium describe a resource based on whole-genome sequencing of 1,135 A. thaliana genomes from Europe, North Africa, and North America. This data set provides high-quality genomic data suitable for GWAS and population genetics studies. Resources from this project are available through web-based tools at http://1001genomes.org/tools/, which can be used to answer questions on genetic diversity and environmental adaptations. For example, SNPs correlated with climate factors were identified related to drought response and fungal defense. Furthermore, ancient expansion of glaciers has shaped A. thaliana populations, and most extant A. thaliana populations can be traced to a single population expansion event when species from North Africa colonized Europe and the British Isles. A limited number of relict A. thaliana populations exist in geographic areas, such as the Iberian Peninsula, that have experienced minimal changes in climate since the last glaciers occupied these areas. A major conclusion from analysis of these 1,135 genomes is A. thaliana contains DNA from multiple ancestors, but a single historical population expansion has favored descendants from one geographic area, which is still unknown. (Summary by Daniel Czerny) Cell 10.1016/j.cell.2016.05.063
There are many accessions of Arabidopsis thaliana beyond the ecotypes predominantly used in research laboratories. In this article, The 1001 Genomes Consortium describe a resource based on whole-genome sequencing of 1,135 A. thaliana genomes from Europe, North Africa, and North America. This data set provides high-quality genomic data suitable for GWAS and population genetics studies. Resources from this project are available through web-based tools at http://1001genomes.org/tools/, which can be used to answer questions on genetic diversity and environmental adaptations. For example, SNPs correlated with climate factors were identified related to drought response and fungal defense. Furthermore, ancient expansion of glaciers has shaped A. thaliana populations, and most extant A. thaliana populations can be traced to a single population expansion event when species from North Africa colonized Europe and the British Isles. A limited number of relict A. thaliana populations exist in geographic areas, such as the Iberian Peninsula, that have experienced minimal changes in climate since the last glaciers occupied these areas. A major conclusion from analysis of these 1,135 genomes is A. thaliana contains DNA from multiple ancestors, but a single historical population expansion has favored descendants from one geographic area, which is still unknown. (Summary by Daniel Czerny) Cell 10.1016/j.cell.2016.05.063








Leave a Reply
Want to join the discussion?Feel free to contribute!