UMP pyrophosphorylase-a moonlighting protein with essential functions in chloroplast development and photosynthesis establishment
Pyrimidine nucleotides (e.g., dCTP, dTTP, UTP, and CTP) are essential building blocks for DNA and RNA synthesis in all organisms. In plants, sucrose and cell wall polymer synthesis depend on pyrimidine nucleotide-derived substrates, such as UDP-glucose, and important classes of membrane lipids are made using CDP-activated substrates. In addition to de novo synthesis pathways, pyrimidine nucleotides are produced via salvage pathways. Uracil is salvaged by uracil phosphoribosyl transferase (UPRT; EC 2.4.2.9), using phosphoribosyl pyrophosphate as substrate to produce UMP, while uridine is salvaged by uridine kinase (UK; EC 2.7.1.48), which uses ATP to phosphorylate uridine to UMP. In this manuscript (Ohler et al., 2019), the authors show that pyrimidine salvage in Arabidopsis occurs predominantly via uridine/cytidine (see figure), with the main enzymes being the bifunctional cytosolic uridine/cytidine kinases 1 and 2 (UCK1 and UCK2; EC 2.7.1.48). A double knockout of UCK1 and UCK2 leads to severe growth restrictions in green tissue (Chen and Thelen, 2011), indicating that Arabidopsis relies on both de novo pyrimidine synthesis and pyrimidine salvage via cytosolic uridine/cytidine. These enzymes have a UPRT domain but do not exhibit any UPRT activity. The only enzyme responsible for UPRT activity in Arabidopsis is the chloroplastic UMP pyrophosphorylase (UPP; EC 2.4.2.9), which is involved in pyrimidine salvage via uracil, but despite being functional, this plastidic uracil salvage pathway has a non-essential role in Arabidopsis.
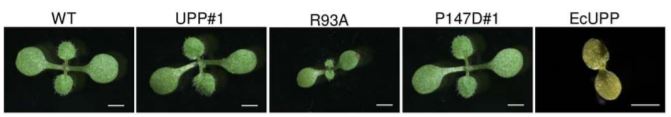
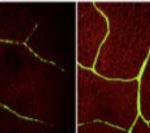
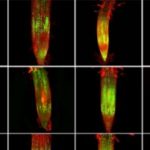
Surprisingly, upp-1, a UPP T-DNA insertion line, germinates but dies at the seedling stage. This knockout exhibits severe growth defects, with pale yellow leaves, reduced chloroplast size and lack of starch synthesis, pointing to a role for UPP in chloroplast development and photosynthesis establishment. Gene expression analysis of young upp-1 seedlings showed no significant changes in the regulation of chloroplast-encoded genes. However, the seedlings had reduced transcript levels for several nuclear-encoded components of photosystem II: two light harvesting chlorophyll binding proteins (LHCB1.4 and LHCB2.3), and a “Photosystem II 5 kDa protein”.
The light-dependent expression of UPP shown by the authors and the wild-type-like growth of upp-1 in darkness, shown by Mainguet et al. (2009), further support a function for UPP in photosynthesis establishment. However, it was an open question whether the UPRT catalytic activity of UPP or a secondary, non-catalytic function (i.e. a moonlighting function) is essential for photosynthesis establishment. Addressing this question, the authors showed by genetic complementation experiments that the lethal phenotype of upp-1 could not only be rescued by expression of wild-type UPP under the control of the UBQ10 promoter, but also by expression of mutated UPP proteins with almost no UPRT activity, generated by substitution of amino acids that are critical for catalysis. However, expression of a catalytically active UPP from a non-photosynthetic organism, Escherichia coli, in the Arabidopsis upp-1 did not rescue the phenotype. These results indicate that UPP plays a vital role in the establishment of photosynthesis, but this is independent of the protein’s UPRT activity. Interestingly, the two enzymes downstream of UPP in the plastidic uracil salvage pathway, UMP kinase (PUMPKIN; EC 2.7.1.14) and nucleoside diphosphate kinase 2 (NDPK2; EC 2.7.4.6), are also moonlighting proteins that function in photosynthetic mechanisms (Schmidt et al., 2019; Dorion and Rivoal, 2015). These findings not only reveal a crucial role for UPP in establishing photosynthetic capacity, but also add to a growing list of moonlighting plant enzymes that have dual catalytic and non-catalytic functions (Claeys et al., 2019).
Figure legend: Overview of pyrimidine de novo synthesis and pyrimidine salvage pathways in Arabidopsis. Pyrimidine de novo synthesis de novo pathway is distributed between chloroplast and cytosol (dashed line). Pyrimidine salvage occurs predominantly via uridine/cytidine, while salvage via uracil has a non-essential role (in grey). UCK1 and UCK2 (uridine/cytidine kinases 1 and 2; EC 2.7.1.48), CDA1 (cytidine deaminase 1; EC 3.5.4.5), NSH1 (nucleoside hydrolase 1; EC 3.2.2.8), Pluto (plastidic nucleobase transporter), UPP (UMP pyrophosphorylase, EC 2.4.2.9), PUMPKIN (UMP kinase; EC 2.7.1.14), NDPK2 (diphosphate kinase 2; EC 2.7.4.6).
LITERATURE CITED
Chen, M. and Thelen, J.J. (2011) Plastid uridine salvage activity is required for photoassimilate allocation and partitioning in Arabidopsis. Plant Cell, 23, 2991-3006.
Claeys, H., Vi,S.L., Xu, X., Satoh-Nagasawa, N., L. Eveland, A.L., Goldshmidt, A., Feil, R., Beggs, G.A., Sakai, H., Brennan, R. G., Lunn, J.E. and Jackson, D. (2019) Control of meristem determinacy by trehalose 6-phosphate phosphatases is uncoupled from enzymatic activity. Nature Plants, Brief Communication, Published: 01 April 2019.
Dorion, S. and Rivoal, J. (2015). Clues to the functions of plant NDPK isoforms. Naunyn-Schmiedeberg’s archives of pharmacology 388:119-132.
Mainguet, S.E., Gakiere, B., Majira, A., Pelletier, S., Bringel, F., Guerard, F., Caboche, M., Berthome, R. and Renou, J.P. (2009) Uracil salvage is necessary for early Arabidopsis development. Plant Journal, 60, 280-291.
Ohler, L., Niopek-Witz S., Mainguet S.E., Möhlmann T. (2019) Physiological function of pyrimidine salvage and interaction with chloroplast biogenesis. Accepted: 01 May 2019.
Schmid, L.M., Ohler, L., Möhlmann, T., Brachmann, A., Muiño, J. M., Leister, D., Meurer, J. and Manavski, N. (2019) PUMPKIN, the sole plastid UMP kinase, associates with group II introns and alters their metabolism. Plant Physiol. 179: 248-264.