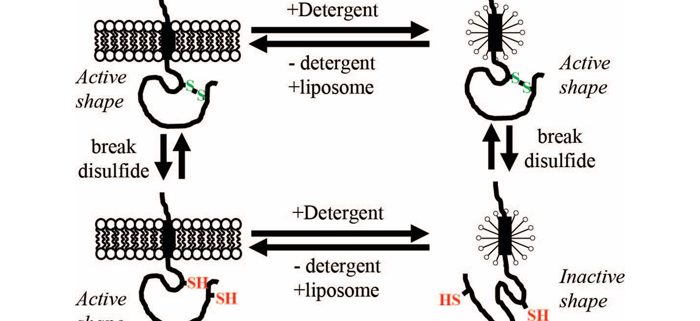
Thylakoid membranes or a disulfide bond help to properly fold Plsp1, a protein that is essential for photosynthesis
McKinnon et al. investigate the effects of a membrane on the folding of a thylakoid protease. Plant Cell https://doi.org/10.1105/tpc.19.00797
By Lucas McKinnon and Steven Theg
Background: Inside a chloroplast is the light-absorbing membrane called the thylakoid, which contains many different proteins and lipid molecules needed for photosynthesis. Physical interactions with membrane lipids are critical for membrane proteins to maintain their 3D shape. Maintenance of the right shape can also be assisted by the formation of disulfide bonds, and this could be especially important if a protein exists in a rapidly changing molecular environment. Lipids and disulfide bonds likely influence the shape and function of Plsp1, an essential enzyme in thylakoids that cleaves off the targeting signal from many thylakoid proteins. An isolated form of Plsp1 has a disulfide bond that, when broken, causes the protein to become inactive likely because of a shape change.
Question: We wanted to study the influence of other proteins and/or thylakoid lipids on the apparent shape change in Plsp1. We used biochemistry tools on Plsp1 isolated in detergents and molecular biology approaches with Plsp1 inside chloroplasts.
Findings: Isolated Plsp1 changes its shape when its disulfide bond is broken, but this shape change does not occur when the protein is inside chloroplasts from Arabidopsis or pea plants. Instead of interactions with other thylakoid proteins, interactions with membrane lipids were observed to be responsible for this phenomenon. A mutant version of Plsp1 that cannot form the disulfide bond went from an inactive to an active form simply by being re-inserted into a liposome (i.e., lab-made lipid bilayer). This re-activated protein shifted back to the inactive form when the pH was reduced, a condition that Plsp1 experiences when thylakoids absorb light. We conclude that the lipid component of the thylakoid membrane act as a protein-folding chaperone towards Plsp1 and that the disulfide bond maintains the active shape of the protein under naturally occurring destabilizing conditions.
Next steps: An important next step in this study is to uncover the mechanism of membrane chaperoning of Plsp1, including elucidating whether specific lipid molecules are involved. Another critical question is related to the exact condition(s) that make the disulfide bond in Plsp1 necessary to maintain the shape and function of the protein in living plants.
Lucas J. McKinnon, Jeremy Fukushima, Joshua K. Endow, Kentaro Inoue and Steven M. Theg. (2020). Membrane chaperoning of a thylakoid protease whose structural stability is modified by the proton motive force. Plant Cell. https://doi.org/10.1105/tpc.19.00797.