The Plant Cell Reviews Plant Immunity: Receptor-Like Kinases, ROS-RLK Crosstalk, Quantitative Resistance, and the Growth/Defense Tradeoff
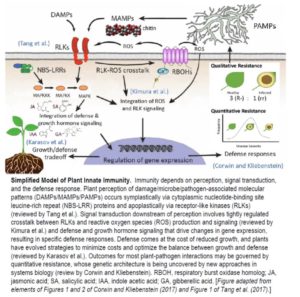 Tender green leaves and tasty tubers, roots, and stems are vulnerable to a wide range of pathogens, pests, and herbivores. Perhaps it should not be surprising that they have evolved an equally wide range of defense mechanisms. This issue of The Plant Cell includes reviews of just a few of the many facets of plant immunity.
Tender green leaves and tasty tubers, roots, and stems are vulnerable to a wide range of pathogens, pests, and herbivores. Perhaps it should not be surprising that they have evolved an equally wide range of defense mechanisms. This issue of The Plant Cell includes reviews of just a few of the many facets of plant immunity.
Plant receptor-like kinases (RLKs) comprise a superfamily of transmembrane proteins, many of which function in pathogen detection as Pattern Recognition Receptors (PRRs). Tang et al. (2017) review recent work on plant RLKs, with a focus on plant-pathogen interactions. The authors present an extended list of known PRRs involved in plant immunity, review their mechanism of action (ligand binding and the formation and regulation of PRR receptor complexes), and highlight new approaches for discovering microbial patterns and their cognate plant PRRs. RLKs play important roles in growth/development as well as immunity, and the authors review what is known about crosstalk and coordination of these pathways. For example, some pathogen effectors actively manipulate RLKs to increase plant susceptibility by mimicking host plant peptide hormones.
Reactive oxygen species (ROS) are produced in response to pathogen invasion and other environmental signals in a manner that is intimately tied to RLK signaling. Kimura et al. (2017) review the role of ROS in RLK signaling and how the regulation of ROS-RLK crosstalk may be a critical component of communication between the cell interior and the environment. The authors discuss the importance of the apoplast as an arena for ROS production and the initiation of ROS-RLK crosstalk, the role of oxidative post-translational modifications in ROS perception, and propose a role for RLK-dependent modulation of both apoplastic and intracellular conditions in facilitating ROS perception and signaling. The importance of RLK-ROS crosstalk is illustrated in the context of stomatal immunity, where the control of stomatal closure restricts a pathogens entry into the plant. Stomatal closure is controlled by a complex interplay of signaling pathways dependent on abscisic acid and other plant hormones, which also intersect with immunity signaling pathways that involve numerous RLK- and ROS-dependent events. Although guard cell ROS production is poorly understood, RLKs appear to play a key role.
Traditional approaches to understanding plant immunity often focus on individual resistance genes that have large effects on plant disease resistance in specific contexts. Corwin and Kliebenstein (2017) argue that quantitative resistance governs most plant-pathogen interactions by the combined effects of many genes with small to moderate effects. The complexity of quantitative resistance and limited power of mapping populations used to study the trait have constrained efforts to understand the underlying mechanisms. The development of new mapping populations with improved power and resolution in a number of model species, such as maize and rice in addition to Arabidopsis, combined with systems biology approaches, such as whole-transcriptome gene expression QTL analysis, are helping to identify causal loci and understand the molecular basis of quantitative resistance. Broad-spectrum resistance is an important concept in plant immunity, but the authors argue that it is poorly understood, and the classical definition of the ability of a host plant to resist multiple genotypes within a single pathogen species may be evolutionarily distinct from the ability to resist specific isolates from numerous different pathogen species. Thus, an important element of studies aimed at understanding broad spectrum resistance may be to consider the role of host and pathogen variation in quantitative resistance across pathogen species.
Plant defense should theoretically impart a cost of reduced growth and reproduction as resources must be diverted to ward off an attack. Karasov et al. (2017) review various means by which plants are able to minimize these costs and optimize growth and defense. Tight regulation of antagonistic crosstalk among growth and defense signaling pathways has been well-characterized, but the selective advantages and evolution of such crosstalk is poorly understood. The authors discuss three types of defense regulation that occur over different time-frames in the plant life cycle: directly induced responses, priming, and transgenerational defense induction. Directly induced responses act to suppress the effects of an ongoing infection and avoid the cost of constitutive expression of defense; priming allows for faster induction of defense to future attacks after the establishment of resistance to an initial attack, and transgenerational memory sets up primed or constitutive means of fighting infection in a future generation. A case study of R genes highlights how evolution may have shaped genetic architecture to allow fine-scale regulation of defense genes to optimize the defense/growth tradeoff. Finally, the authors discuss new evidence that plants exert genetic control over their microbiome to optimize defense.
These four reviews covering different but highly related aspects of plant immunity provide excellent background, explore exciting new directions, and offer insightful perspectives to stimulate future research in this important field of plant biology.
IN BRIEF by Nancy Eckardt neckardt@aspb.org
REFERENCES
Corwin, J.A., and Kliebenstein, D.J. (2017). Quantitative Resistance: More than Just Perception of a Pathogen. Plant Cell doi: 10.1105/tpc.16.00915.
Karasov, T.L., Chae, E., Herman, J.J., and Bergelson, J. (2017). Mechanisms to mitigate the growth/defense tradeoff. Plant Cell doi: 10.1105/tpc.16.00931.
Kimura, S., Waszczak, C. Hunter, K., and Wrzaczek, M. (2017). Bound by Fate: The Role of Reactive Oxygen Species in Receptor-Like Kinase Signaling. Plant Cell doi: 10.1105/tpc.16.00947.
Tang, D., Wang, G., and Zhou, J.-M. (2017). Receptor kinases in plant-pathogen interactions: more than pattern-recognition. Plant Cell doi: 10.1105/tpc.16.00891.





Leave a Reply
Want to join the discussion?Feel free to contribute!